- Home
- /
- Programming
- /
- SAS Procedures
- /
- Welch's ANOVA inconsistent in PROC MIXED vs. PROC GLM
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I'm trying to implement Welch's ANOVA in SAS, and, according to the documentation at http://support.sas.com/kb/22/526.html, I'm supposed to be able to do it in two equivalent ways:
PROC GLM;
CLASS trt;
MODEL y = trt;
MEANS trt / WELCH;
RUN;
or
PROC MIXED;
CLASS trt;
MODEL y = trt / DDFM=SATTHERTH;
REPEATED / GROUP=TRT;
LSMEANS trt;
RUN;
However, I only get the same results (*F*-statistic and *p*-value) from both when I have two groups, but not more than two (six, specifically).
Online, I've found examples of others also using both procedures for the "same" Welch's ANOVA, but their results also differ when I run them. Are the d.f. actually approximated slightly differently in both procedures?
Here is a working example: It's the hermit crab data from Design of Experiments: Statistical Principles of Research Design and Analysis (2nd Ed.) by Kuehl.
DATA
DATA hermits;
DO coastal_site=1 to 6;
DO rep=1 to 25;
INPUT hermit_count @@;
OUTPUT;
END;
END;
DATALINES;
0 0 22 3 17 0 0 7 11 11 73 33 0 65 13 44 20 27 48 104 233 81 22 9 2
415 466 6 14 12 0 3 1 16 55 142 10 2 145 6 4 5 124 24 204 0 0 56 0 8
0 0 4 13 5 1 1 4 4 36 407 0 0 18 4 14 0 24 52 314 245 107 5 6 2
0 0 0 4 2 2 5 4 2 1 0 12 1 30 0 3 28 2 21 8 82 12 10 2 0
0 1 1 2 2 1 2 29 2 2 0 13 0 19 1 3 26 30 5 4 94 1 9 3 0
0 0 0 2 3 0 0 4 0 5 4 22 0 64 4 4 43 3 16 19 95 6 22 0 0
;
RUN;
PROC GLM version of Welch's ANOVA
PROC GLM DATA=hermits;
CLASS coastal_site;
MODEL hermit_count = coastal_site;
MEANS coastal_site / WELCH;
RUN;
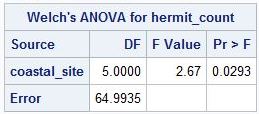
PROC GLM output
PROC MIXED version of Welch's ANOVA
PROC MIXED DATA=hermits;
CLASS coastal_site;
MODEL hermit_count=coastal_site / DDFM = SATTERTHWAITE;
REPEATED / GROUP = coastal_site;
LSMEANS coastal_site;
RUN;
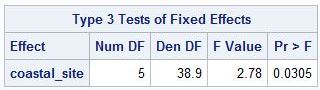
PROC MIXED output
Can anyone tell me why these are not giving the same test statistics? If they, in general, don't give the same answers (beyond two groups), which one should be trusted (i.e., which one is implementing the test correctly)? Is it just a matter of slightly different
[1]: http://i.stack.imgur.com/bmDHI.jpg
[2]: http://i.stack.imgur.com/hchzJ.jpg
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I am going to go off topic here, as answering the question posed really won't get you any closer to the right answer. Those count data are definitely not going to result in normally distributed errors, and consequently neither PROC GLM nor PROC MIXED is appropriate for the analysis.
Try this for a heterogeneous variance with overdispersion analysis:
PROC GLIMMIX DATA=hermits;
CLASS coastal_site;
MODEL hermit_count = coastal_site/dist=poisson ddfm=satterthwaite;
LSMEANS coastal_site / diff ilink;
RANDOM _residual_/group=coastal_site;
RUN;
Steve Denham
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks, Steve. Yes, I realize these data are not meeting assumptions, but was using it as a minimal working example since I had it readily available. Despite not meeting the assumptions, if both PROC GLM and PROC MIXED use the same theory for Welch's test, they should give the same results on the same set of data.
In actuality, once the data are log-transformed, unequal variances aren't really a problem, but, again, I would like to know why these two procedures give different results.
Others may find your suggested use of PROC GLIMMIX useful in general for count data, however, so thanks for sharing.
Thank you,
Megan
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Now it gets interesting. Try running your MIXED code with ddfm=kenwardrogers. My simulated data now puts the denominator degrees of freedom much closer to that from Welch's ANOVA. Since the Kenward-Rogers method depends on applying shrinkage before calculating Satterthwaite degrees of freedom, I feel more comfortable using it in PROC MIXED than just the ddfm=satterthwaite. However, it has been known to result in values for numerator degrees of freedom that don't make a lot of sense.
Still, for data with heterogeneous variances, I prefer the REML of MIXED to the sums of squares of GLM, and would apply the Kenward-Rogers method, rather than Satterthwaite. This would be even a stronger preference in the face of unbalanced data, or repeated/random effects.
Maybe someone can jump in with matrix math to show why the results are identical at m=2 groups, and diverge for m>2.
Steve Denham
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks again, Steve. I was thinking Welch's probably uses different d.f. I just find it interesting that the SAS documentation doesn't make mention of the fact that these two sets of code they present (that they say give the same test) are not actually equivalent (or define when they would be). I was going to try to work out the math at some point, but just do not have the time (90 hours/week is good enough for me right now...).
I also prefer PROC MIXED, which is why I was curious about this. The PROC GLM code with WELCH option kind of makes you "feel" like this must be the "better" code since it says WELCH right in it, but obviously this isn't sound logic.
My biggest qualm is the lack of documentation, even though my guess is the results are usually quite "close" (if we're willing to "accept" that as "good enough").
I've since found this info on Cross Validated, which I think may be what's going on, and leads me to favor using PROC MIXED with DDFM=SATTERTHWAITE over PROC GLM with WELCH (comments are also useful): http://stats.stackexchange.com/questions/70959/should-i-use-welchs-1947-approximate-degrees-of-freed...
Maybe I'll fill out a feedback form asking for SAS to clarify their page from the original post...
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →Learn the difference between classical and Bayesian statistical approaches and see a few PROC examples to perform Bayesian analysis in this video.
Find more tutorials on the SAS Users YouTube channel.
SAS Training: Just a Click Away
Ready to level-up your skills? Choose your own adventure.