- Home
- /
- Programming
- /
- Graphics
- /
- Errant graph when plotting predicted values from proc nlin
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hello,
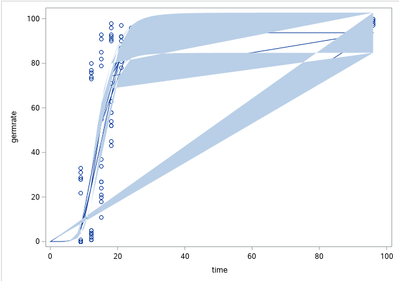
I'm currently working on a SAS Programm to fit my data of a germinating fungus to a given logistic function. This works fine for now, as the estimated values fit well to the data and predicted values based on that function seem acceptable. But when I try to plot this data the resulting graph looks like this:
Does anyone has any suggestions regarding this problem? I'm using SAS University Edition / SAS Studio 3.8
Thanks for your help!
The code used to create this graph is as follows:
data fungi;
input time germrate;
datalines;
9 0
9 0
9 0
9 0
9 0
9 0
9 0
9 0
12 1
12 1
12 1
9 1
12 2
12 3
12 4
12 4
12 5
12 5
15 11
15 18
15 20
15 21
9 22
15 24
15 27
15 27
9 28
9 28
9 29
9 31
9 33
15 34
15 37
18 43
18 45
18 52
18 52
18 54
15 55
18 58
18 58
18 63
18 71
21 72
12 73
21 74
12 74
12 76
12 76
12 77
21 78
15 79
12 80
21 81
15 82
21 83
21 84
21 84
24 85
15 85
24 86
21 86
24 87
21 87
21 87
24 88
24 90
24 90
24 90
21 90
18 90
18 91
15 91
24 92
24 92
21 92
21 92
21 92
18 92
24 93
24 93
18 93
15 93
24 94
24 94
24 95
24 96
18 96
21 97
18 98
96 99
96 100
96 99
96 98
96 97
96 100
;
proc nlin data=fungi;
parms Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
run;
proc nlin data=fungi method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean l95=l95ind u95=u95ind;
run;
proc print data=nlinout;
run;
data filler;
do time = 0 to 96 by 2;
predict=1;
output;
end;
run;
data fitthis;
set fungi filler;
run;
proc nlin data=fitthis method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean;
run;
proc print data=nlinout(where=(predict=1)); run;
proc sgplot data=nlinout;
scatter x=time y=germrate;
series x=time y=pred;
band upper=u95mean lower=l95mean x=time ;
run;
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi @RobinN , you just need to sort your graph by time before you do the plot. Please see the example below.
data fungi;
input time germrate;
datalines;
9 0
9 0
9 0
9 0
9 0
9 0
9 0
9 0
12 1
12 1
12 1
9 1
12 2
12 3
12 4
12 4
12 5
12 5
15 11
15 18
15 20
15 21
9 22
15 24
15 27
15 27
9 28
9 28
9 29
9 31
9 33
15 34
15 37
18 43
18 45
18 52
18 52
18 54
15 55
18 58
18 58
18 63
18 71
21 72
12 73
21 74
12 74
12 76
12 76
12 77
21 78
15 79
12 80
21 81
15 82
21 83
21 84
21 84
24 85
15 85
24 86
21 86
24 87
21 87
21 87
24 88
24 90
24 90
24 90
21 90
18 90
18 91
15 91
24 92
24 92
21 92
21 92
21 92
18 92
24 93
24 93
18 93
15 93
24 94
24 94
24 95
24 96
18 96
21 97
18 98
96 99
96 100
96 99
96 98
96 97
96 100
;
proc nlin data=fungi;
parms Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
run;
proc nlin data=fungi method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean l95=l95ind u95=u95ind;
run;
proc print data=nlinout;
run;
data filler;
do time = 0 to 96 by 2;
predict=1;
output;
end;
run;
data fitthis;
set fungi filler;
run;
proc nlin data=fitthis method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean;
run;
proc print data=nlinout(where=(predict=1)); run;
proc sort data = nlinout;
by time;
run;
proc sgplot data=nlinout;
scatter x=time y=germrate;
series x=time y=pred;
band upper=u95mean lower=l95mean x=time ;
run;- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi @RobinN , you just need to sort your graph by time before you do the plot. Please see the example below.
data fungi;
input time germrate;
datalines;
9 0
9 0
9 0
9 0
9 0
9 0
9 0
9 0
12 1
12 1
12 1
9 1
12 2
12 3
12 4
12 4
12 5
12 5
15 11
15 18
15 20
15 21
9 22
15 24
15 27
15 27
9 28
9 28
9 29
9 31
9 33
15 34
15 37
18 43
18 45
18 52
18 52
18 54
15 55
18 58
18 58
18 63
18 71
21 72
12 73
21 74
12 74
12 76
12 76
12 77
21 78
15 79
12 80
21 81
15 82
21 83
21 84
21 84
24 85
15 85
24 86
21 86
24 87
21 87
21 87
24 88
24 90
24 90
24 90
21 90
18 90
18 91
15 91
24 92
24 92
21 92
21 92
21 92
18 92
24 93
24 93
18 93
15 93
24 94
24 94
24 95
24 96
18 96
21 97
18 98
96 99
96 100
96 99
96 98
96 97
96 100
;
proc nlin data=fungi;
parms Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
run;
proc nlin data=fungi method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean l95=l95ind u95=u95ind;
run;
proc print data=nlinout;
run;
data filler;
do time = 0 to 96 by 2;
predict=1;
output;
end;
run;
data fitthis;
set fungi filler;
run;
proc nlin data=fitthis method=newton;
parameters Pmax=98 teta=15 ;d=6 ;
model germrate = Pmax*(1-(1/(1+((time/teta)**d))));
output out=nlinout predicted=pred l95m=l95mean u95m=u95mean;
run;
proc print data=nlinout(where=(predict=1)); run;
proc sort data = nlinout;
by time;
run;
proc sgplot data=nlinout;
scatter x=time y=germrate;
series x=time y=pred;
band upper=u95mean lower=l95mean x=time ;
run;- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →Learn how use the CAT functions in SAS to join values from multiple variables into a single value.
Find more tutorials on the SAS Users YouTube channel.
SAS Training: Just a Click Away
Ready to level-up your skills? Choose your own adventure.