- Home
- /
- Analytics
- /
- Stat Procs
- /
- Logistic regression Analysis
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I have a new request to create a statistics table with logistic regression. I have attached a sample with this for reference
Expect Somme insights on this since Iam new to this scenario
Covariate Estimate odds ratio 95%ci waldchisqr pvalue
These needs to find
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
proc logistic data=sashelp.heart;
class sex bp_status;
model status=sex bp_status weight height/clparm=wald clodds=wald ;
/*oddsratio sex/cl=wald;*/
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi I have found a sample code, but i needed the below results for my requiremnt. I am not understanding how to pass the proper variables and options . the requirement below
| Covariate | Estimate | Odds Ratio | 95% CI | Wald Chi-Square | P-value |
| Results | xx.xx | xx.xx | xx.xx | xx.xx | xx.xx |
| Male | xx.xx | xx.xx | xx.xx | xx.xx | xx.xx |
| Female | Reference | Reference | Reference | Reference | Reference |
| Results*Male | xx.xx | xx.xx | xx.xx | xx.xx | xx.xx |
| Results*Female | Reference | Reference | Reference | Reference | Reference |
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This is basic output from PROC LOGISTIC. But what you want appears in two separate tables. Below assumes that your binary response is called Response, and has values 0 or 1 with 1 being the event level of interest.
proc logistic data=YourDataSet;
class gender(ref='Male');
model Response(event="1")=Results gender Results*gender / clparm=wald;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi I have used the code,.. the problem is that I am not able to get the ODDS ratio
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
@ambadi007 wrote:
Hi I have used the code,.. the problem is that I am not able to get the ODDS ratio
Please show us your code. Please show us the output. Please show us (a portion) of your data.
Paige Miller
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Please find the data and the code below which i have.
ods trace on;
ods output Association=or2 GlobalTests=GBLT_2;
proc logistic data=mydata outmodel=stat_3;
class gender (ref="F");
model improvement(event="Y")=result gender gender*result / clodds=wald;
run;
ods trace off;
| improvement | Result | Gender |
| Y | 12 | M |
| N | 13 | F |
| Y | 14 | M |
| Y | 15 | F |
| Y | 16 | M |
| Y | 17 | M |
| Y | 18 | F |
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Please, in the future, follow Maxim 2, which says:
Maxim 2
Read the log.
Everything you need to know about your program is in the log. Interpreting messages and NOTEs is essential in finding errors.
The log says:
NOTE: The CLODDS= option does not compute odds ratios for effects involved in interactions or nestings.
Specify the ODDSRATIO statement to compute odds ratios for these effects.
So you need to add the ODDSRATIO command into your code. Maybe like this:
proc logistic data=mydata outmodel=stat_3;
class gender (ref="F");
model improvement(event="Y")=result gender gender*result / clodds=wald;
oddsratio gender;
oddsratio result;
run;
The log also says
WARNING: There is a complete separation of data points. The maximum likelihood estimate does not exist.
WARNING: The LOGISTIC procedure continues in spite of the above warning. Results shown are based on the last maximum likelihood iteration. Validity
of the model fit is questionable.
which means that the results you get are questionable.
Paige Miller
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
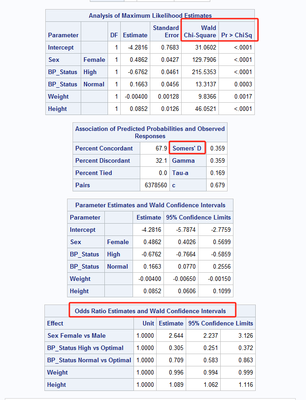
proc logistic data=sashelp.heart(obs=1000);
class sex bp_status;
model status=sex bp_status weight height sex*height/clparm=wald clodds=wald ;
oddsratio height/at(sex=all) cl=wald;
run;
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.