- Home
- /
- Analytics
- /
- Stat Procs
- /
- Bizarre residuals PROC MIXED
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
EDIT: I now found that if I omit initial plant height ("initht") in the random statement, I do not have the issue with the residuals described below. Hopefully, this can lend some insight.
---------------
Hi all,
I am fitting a model with PROC MIXED.
I have a continuous response variable "biomass" and 3 predictor variables 1) light (2 level categorical) 2) flood depth (ranked ordinal) and 3) flood duration (ranked ordinal), and I am fitting all 2 and 3 way interactions.
I have 2 random effects, initial plant height and experimental block.
Here is my model statement (flood depth is "sublevel").
PROC MIXED data=RUN1biomass covtest plots=(residualpanel);
CLASS Block Light sublevel duration;
MODEL AGB=Light Sublevel duration
duration*Light
SubLevel*Light SubLevel*Duration Sublevel*Light*Duration / influence DDFM=SATTERTH OUTP=work.diagnostics RESIDUAL;
RANDOM Block initht;
run;
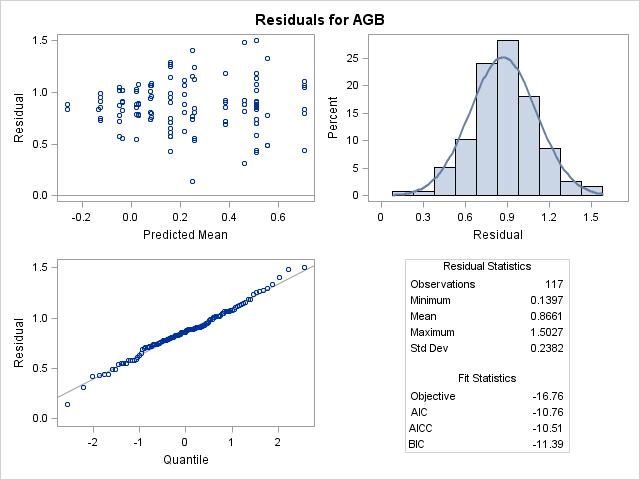
The model runs fine but I don't understand the residuals. I am attaching the residual panel produced by proc mixed. Notice that ALL THE RESIDUALS ARE POSITIVE. How is this even possible?? Is this some glitch or is there any way this actually makes some sense? The studentized residual panels and pearson residual plots look basically the same. All of the conditional residual plots look good--normal distribution of residuals and random scatter around 0.
Thank you!!
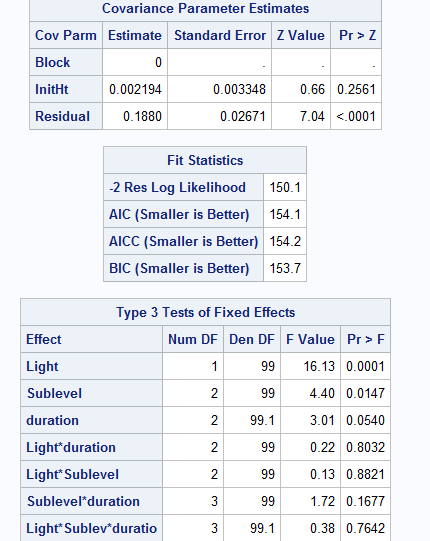
And here is the output from proc mixed
- Tags:
- proc mixed
- residuals
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I don't see there is an issue from the residual plot. The horizontal axis in the residual histogram is from 0.0 to 1.5 and is normally distributed. This indicates all the predictions are larger than the raw data, and therefore the residuals are positive.
One thing is more serious from this output is that the model might be overfitting. Because the covariance parameter estimates for effect block is 0. You should see some notes in the log window such as "NOTE : Estimated G matrix is not positive definite ". This means there is not much variation between blocks. You might consider take out effect BLOCK and rerun the model to see what happens.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You are right: residuals that do not have mean = 0 indicate a serious problem with the model. Good spotting! As you note in your update, the fundamental problem was specifying INITHT as a random effects factor. But there may be other issues.
You have not provided enough information to determine the correct model. Let's say that you have blocks, and within each block you have multiple plots. BLOCKs and PLOTs comprise random effects factors. Each fixed effects factor (LIGHT, SUBLEVEL, and DURATION) is associated (ideally, randomly assigned) to a level of one of the random effects factors. It could be that all three fixed effects factors are assigned to individual PLOTs. Or it could be that one (or more) fixed effects factors are assigned to BLOCKs, and the others are assigned to PLOTs. The statistical model must reflect the experimental design.
Note that ANOVA (where LIGHT, SUBLEVEL, and DURATION are classification effects) does not take into account whether the levels of a fixed effects factor are ordered. To accommodate ordered levels, you have to either regress on the fixed effects factor (which can no longer be listed in the CLASS statement) or specify contrasts (the LSMESTIMATE statement is very handy) that address the ordered levels (i.e., linear, quadratic, etc.).
I presume that you have appreciable variability among PLANTs in INITHT and that there is a relationship between AGB and INITHT that you would like to incorporate into the statistical model. One approach is to use an analysis of covariance model (ANCOVA) in which INITHT is a continuous-scale fixed effects factor (and so is included in the MODEL statement, but not in the CLASS statement, because the model is regressing AGB on INITHT) and which potentially interacts with other fixed effects factors (LIGHT, SUBLEVEL, and DURATION). Depending on the experimental design, the statistical model may be a random coefficients model (essentially a regression in a mixed model); but we don't have enough information to tell.
This brings us to PLANT, which is the unit on which INITHT, and presumably AGB, is measured. Is PLANT equivalent to PLOT, or are there multiple PLANTs within each PLOT? Multiple PLANTs within a PLOT typically would be considered as subsamples. The design structure--whatever it is--must be reflected in the statistical model.
As you might imagine, there are many subtleties to the correct implementation of subsamples, ordinal contrasts, ANCOVA-with-interactions models, and random coefficients models in a statistical model. Probably more than can be adequately addressed in a forum like this. Hopefully my comments provide you with some places to start.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks to a colleague, I think I now understand the Mystery of the Positive Residuals. I think the OP and others already understand the problem, but for the sake of those who are still confused let me try to explain something that initially confused me.
By default, a residual panel shows marginal residuals. The title of a marginal panel plot is "Residuals for Y," which is the title for the panel that the OP attached.
The PLOTS=RESIDUALPANEL option takes suboptions. If you specify
plots=residualpanel(conditional)
then you get a plot of the conditional residuals. The title for a panel plot of the conditional residuals is "Conditional Residuals for Y."
The OP hinted that "all of the conditional residual plots look good", so apparently realized that the marginal residuals were being displayed. However, I (and maybe other readers) did not appreciate the importance of that comment.
Mathematicaly, conditional residuals will have approximately zero mean. However, marginal residuals do not necessarily have mean zero. The marginal residuals are
R_marginal = Y - X*beta = Z*gamma + epsilon
and one of the columns of Z is the continuous variable "initial plant height," which is all postive. Because of other fitting problems (described by others) the fitted coefficients for the model resulted in positive marginal residuals.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Rick, I'm glad you reminded folks about the difference between marginal and conditional residuals. Both have value for different applications. Conditional residuals are labeled as "Conditional residuals" on the graph, but marginal residuals are justed labeled as "Residuals". That can be confusing.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This is a follow-up to your response to residual from proc mixed. The internally conditional residual is usually obtained from model statement as:
Model Y=trt/ddfm=kr outp= cond_resid residual;
The addition of influence option to model statement will return an externally conditional residual which can be obtained through
ods output Influence = output_dataname.
The output from influence has as well internally conditional residual which differed from what one obtained from model statement through cond_resid file. How do we explain this disparity.
Thanks
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The INFLUENCE option produces internally and externally studentized marginal (not conditional) residuals.
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.