- Home
- /
- Programming
- /
- Programming
- /
- Re: How to create binary variable from existing categorical variable a...
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi,
I'm a new user of SAS 9.4 program and I am working with a data of my research. I have this character column which looks like:
Var1
XY_001
XY_021
XY_001
XY_000
XY_015 .......etc
it is around 633 obsrvations and have also 577 missings! All observations are 1210
and if I run a proc freq for this variable the frequencies of the readings are different ranging from 1 to 38 and the missings
Now I want to create a new variable called Var2 I want to make it binary (0/1). for those categories who have 1 as a frequency to be 0
and those categories who have frequency more or equal than 2 to be 1. and the missings as missing
and to call 0: not group
and 1: group
How can I create the program??
thank you
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Let's start by extending the PROC FREQ that you ran, so it also creates an output data set:
proc freq data=have;
tables var1 / out=counts (keep=var1 count);
run;
Next, sort your original data set:
proc sort data=have;
by var1;
run;
Finally, merge the PROC FREQ results with your data:
data want;
merge have
counts (where=(var1 > ' '));
by var1;
if count=1 then flag=0;
else if count > 1 then flag=1;
drop count;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Let's start by extending the PROC FREQ that you ran, so it also creates an output data set:
proc freq data=have;
tables var1 / out=counts (keep=var1 count);
run;
Next, sort your original data set:
proc sort data=have;
by var1;
run;
Finally, merge the PROC FREQ results with your data:
data want;
merge have
counts (where=(var1 > ' '));
by var1;
if count=1 then flag=0;
else if count > 1 then flag=1;
drop count;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi Astounding,
Thank you for your reply I have an issue:
I run this code:
/* to create new cluster status variable*/
proc freq data=genotyped_1210;
tables clustername / out=counts (keep=clustername count);
run;
/*Next, sort your original data set:*/
Proc sort data= genotyped_1210;
by clustername;
run;
The 1st part wet well, the sorting part gave me this error:
80 Proc sort data= genotyped_1210;
81 by clustername;
82 run;
ERROR: You cannot open WORK.GENOTYPED_1210.DATA for output access with member-level control because
WORK.GENOTYPED_1210.DATA is in use by you in resource environment SORT.
NOTE: The SAS System stopped processing this step because of errors.
NOTE: PROCEDURE SORT used (Total process time):
real time 0.00 seconds
cpu time 0.00 seconds
What do you suggest?
Thank you
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You probably have the original data set open in a data viewer (EG? Check Tools->View Open Data Sets.)
To avoid, you can work from a copy by sorting the original into another file and work forward from there -- or else just make sure all data grid views are closed before running the next step.
/* to create new cluster status variable*/
proc freq data=genotyped_1210;
tables clustername / out=counts (keep=clustername count);
run;
/*Next, sort your original data set:*/
Proc sort data= genotyped_1210 out=sorted_genotyped;
by clustername;
run;- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you ChrisHemedinger,
I closed all the data viewer and it worked!
However, when doing the next program as follows
/*Finally, merge the PROC FREQ results with your data:*/
data TB;
merge sorted_genotyped
counts (where=(clustername > ' '));
by clustername;
if count=1 then unique=0;
else if count > 1 then cluster=1;
drop count;
run;The following error was:
58 data TB;
59 merge sorted_genotyped
60 counts (where=(clustername > ' '));
ERROR: The value is not a valid SAS name.
61 by clustername;
62 if count=1 then unique=0;
63 else if count > 1 then cluster=1;
64 drop count;
65 run;
NOTE: The SAS System stopped processing this step because of errors.
WARNING: The data set WORK.TB may be incomplete. When this step was stopped there were 0
observations and 239 variables.
WARNING: Data set WORK.TB was not replaced because this step was stopped.
NOTE: DATA statement used (Total process time):
real time 0.01 seconds
cpu time 0.01 seconds
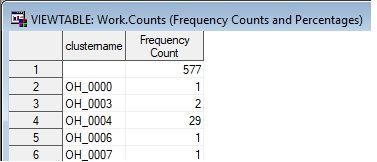
attached picture of example of the character values of the variable: clustername
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This won't explain the error, but it is a small required fix. You now have:
if count=1 then unique=0;
else if count > 1 then cluster=1;
That needs to become:
if count=1 then unique=0;
else if count > 1 then unique=1;
The error message itself it not really explainable. You can try changing the WHERE condition from this:
(where=(clustername > ' '))
to this:
(where=(clustername ne ' '))
Also make sure the quotes around the blank space are really quotes ... you can try deleting them, then typing them back in again. It shouldn't make a difference, but you never know.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks all for your help
The problem still didn't solve it could be because I have an ID numbers in the original file thatcouldaffect the merge step ?!
I will try to use R instead to see if this could solve the problem!
I appreciatr all your suggestions and programs you provided I tried them all and it didn't work.
Thank you so much
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
What is it that you want as your output?
data have ;
length Var1 $10;
input Var1 @@ ;
cards;
XY_001 XY_021 . XY_001 . XY_000 . XY_015 .
;
proc sql noprint ;
create table want as
select var1,count(*) as nobs
, (calculated nobs > 1) as var2
from have
group by 1
;
quit;
proc print; run;Obs Var1 nobs var2 1 4 1 2 XY_000 1 0 3 XY_001 2 1 4 XY_015 1 0 5 XY_021 1 0
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you for your all contributions the problem is solved by R!
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
i have three categorical variables in my data set:
1. alcohol (current=1, other=0)
2. smk (current=1, other=0)
3.drug (current=1, other=0)
now i want to creat a variable that only includes level1 of these variables:
here is what I wrote:
if alcohol=1 & smk=1 & drug=1 then (new variable name)= 1; else (new variable)=0;
however this is not showing in my proc freq.
any advice is appreciated.
Thanks
Catch up on SAS Innovate 2026
Dive into keynotes, announcements and breakthroughs on demand.
Explore Now →Learn how use the CAT functions in SAS to join values from multiple variables into a single value.
Find more tutorials on the SAS Users YouTube channel.
SAS Training: Just a Click Away
Ready to level-up your skills? Choose your own adventure.