- Home
- /
- Programming
- /
- Graphics
- /
- Plot a by b means using SAS UE
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hello,
I'm trying to run an sgplot procedure using SAS UE. The purpose is to plot a variable by another variable means. I have the procedure I am trying to follow that uses gplot (which does not work for SAS UE). I'm not sure what the equivalent would be when using SAS UE. I appreciate any help regarding this!
Procedure given (cannot be used for SAS UE):
proc gplot data=clinical;
axis1 length=4 in;
axis2 length=6 in;
plot dna*lipid=tpa / vaxis=axis1 haxis=axis2;
symbol1 v=J f=special h=3 i=join color=blue;
symbol2 v=K f=special h=3 i=join color=red;
run;
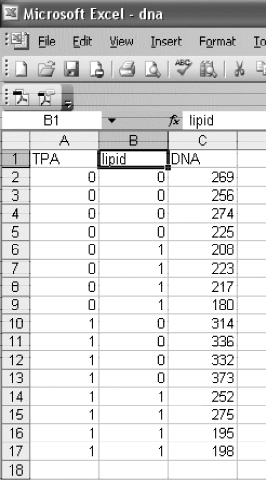
Data:
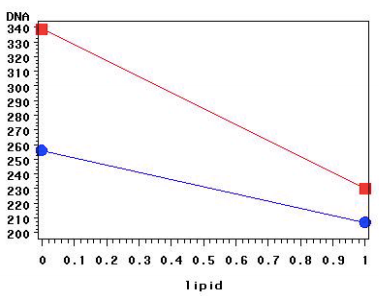
Final plot (tpa by lipid means, goal for SAS UE procedure not yet gotten):
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This is called an "Interaction Plot" and it is produced automatically when you perform a two-way analysis of mean with an interaction term:
data dna;
input TPA lipid DNA;
datalines;
0 0 269
0 0 256
0 0 274
0 0 225
0 1 208
0 1 223
0 1 217
0 1 180
1 0 314
1 0 336
1 0 332
1 0 373
1 1 252
1 1 276
1 1 195
1 1 198
;
proc glm data=DNA plots=IntPlot;
class TPA lipid;
model DNA = TPA | lipid;
run;
If you don't want to see the observed values, you can write the model to an item store and use PROC PLM to create an effect plot:
proc glm data=DNA;
class TPA lipid;
model DNA = TPA | lipid;
store GlmModel;
run;
proc plm restore=GLMModel;
effectplot;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Something like this should do:
proc sql;
create table graph as
select
tpa,
mean(lipid) as lipid,
mean(dna) as dna
from clinical
group by tpa;
quit;
ods graphics / height=4in width=6in;
proc sgplot data=graph noautolegend;
styleattrs
DATACONTRASTCOLORS=(blue red)
DATASYMBOLS=(squarefilled circlefilled);
series y=dna x=lipid / group=tpa
markers markerattrs=(size=8)
lineattrs=(pattern=solid);
run;- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This is called an "Interaction Plot" and it is produced automatically when you perform a two-way analysis of mean with an interaction term:
data dna;
input TPA lipid DNA;
datalines;
0 0 269
0 0 256
0 0 274
0 0 225
0 1 208
0 1 223
0 1 217
0 1 180
1 0 314
1 0 336
1 0 332
1 0 373
1 1 252
1 1 276
1 1 195
1 1 198
;
proc glm data=DNA plots=IntPlot;
class TPA lipid;
model DNA = TPA | lipid;
run;
If you don't want to see the observed values, you can write the model to an item store and use PROC PLM to create an effect plot:
proc glm data=DNA;
class TPA lipid;
model DNA = TPA | lipid;
store GlmModel;
run;
proc plm restore=GLMModel;
effectplot;
run;
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →Learn how use the CAT functions in SAS to join values from multiple variables into a single value.
Find more tutorials on the SAS Users YouTube channel.
SAS Training: Just a Click Away
Ready to level-up your skills? Choose your own adventure.