- Home
- /
- Analytics
- /
- Stat Procs
- /
- Simple main effect code
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hello. I have certain pairs that I am interested in comparing as post-hoc tests after finding significant interaction or main effects of Phoneme_stress.
In the case of significant interaction of Phoneme_stress*Phase_info1, I want to examine:
simple main effects of Phoneme_stress at Phase_info1=1 &
simple main effects of Phoneme_stress at Phase_info1=3.
In the case of insignificant interaction and significant main effect of Phoneme_stress, I want to examine:
Marginal comparison of Phoneme_stress in the following comparisons only:
Phoneme_stress=0 vs. Phoneme_stress=2,
Phoneme_stress=0 vs. Phoneme_stress=3,
Phoneme_stress=0 vs. Phoneme_stress=4,
Phoneme_stress=0 vs. Phoneme_stress=5.
Could you please help me modify below codes to meet my goal?
Thank you in advance.
Linda
proc glimmix data=a1.prediss1 plots=residualpanel(conditional marginal);
class phase_info1 Phoneme_stress subject;
model ResidualError_Sum= phase_info1|Phoneme_stress/solution dist=gamma;
output out=a1.predata pred=pred resid=r;
random intercept phase_info1 Phoneme_stress/ subject=subject;
slice phase_info1*Phoneme_stress / sliceby=Phoneme_stress diff alpha=.0125;
lsmeans Phoneme_stress/diff alpha=.0125;
lsmestimate phase_info1*Phoneme_stress
"Phoneme_stress: 0 v 2" 1 -1 0 0 0;
"Phoneme_stress: 0 v 3" 1 0 -1 0 0;
"Phoneme_stress: 0 v 4" 1 0 0 -1 0;
"Phoneme_stress: 0 v 5" 1 0 0 0 -1;
/ adjust=simulate(seed=29847);
run;
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I am not entirely sure where your confusion lies, but guessing here....
LSMEANS are not the same as MEANS in some situations. See Means versus LS-Means. If you are using descriptive statistics computed from the raw data (using something like the MEANS procedure), then model estimates will not match if the sample sizes differ (i.e., the design is unbalanced).
Descriptive statistics will also differ from model estimates if the response is transformed and the model estimates are subsequently back-transformed to the original scale, or the model is a generalized linear model and model estimates are inverse-linked to the original scale.
Consequently, we need to have a certain amount of trust in the model because the model is often the best source of estimates, and that requires that we have a correctly specified model.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
it's useful to look at the L matrix coefficients when writing out the coefficients for the contrast: https://support.sas.com/documentation/cdl/en/statug/63033/HTML/default/viewer.htm#statug_mixed_sect0...
in glimmix i believe you just use E option on the model statement https://support.sas.com/documentation/cdl/en/statug/63033/HTML/default/viewer.htm#statug_glimmix_a00...
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
How many levels does phase_info1 have?
How many levels does phoneme_stress have?
How many levels are there for the interaction of phase_info1 and phoneme_stress?
Have you read the documentation for the LSMESTIMATE statement, and if the documentation is not quite enough information, have you read CONTRAST and ESTIMATE Statements Made Easy: The LSMESTIMATE Statement?
The order of coefficients in the LSMESTIMATE statement corresponds to the order depicted in the LSMEANS output for phase_info1 * phoneme_stress; the order is determined by the order of factors in the CLASS statement.
You can always check the results of the LSMESTIMATE statement by hand using the pertinent estimates reported by the LSMEANS statements.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you.
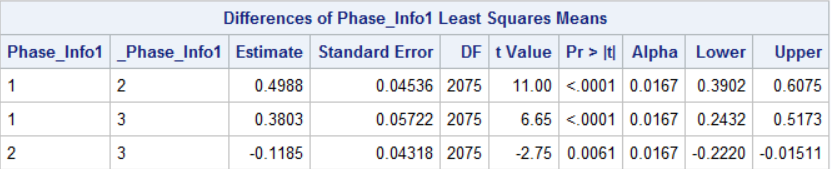
You mentioned that I can calculate the difference by hand, but I thought I could read in the difference from the estimate column.
For example, the DV in phase_info1=1 is higher by 0.4988 than phase_info 1=2.
Was I wrong?
Thanks for your advice in advance.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
My point is that you can check the syntax of any LSMESTIMATE statement by computing its corresponding contrast by hand using the estimates reported by LSMEANS statements. (It's harder to hand-compute the contrast SE, so we let LSMESTIMATE do that for us.)
If the contrast is not complicated--for example, the pairwise comparisons of main effect means for Phase_Info1 in your table--then you do not need to use LSMESTIMATE statements and so you don't need to check whether you've specified the LSMESTIMATE statements correctly.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I see your point.
I came to ask different question, because the estimate value was different from what I obtained as difference value between two means from the descriptive stat table. Now, I have to admit how little I know about all these tools. I have no idea what that estimate is estimating.
Thanks for your advice in advance.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I am not entirely sure where your confusion lies, but guessing here....
LSMEANS are not the same as MEANS in some situations. See Means versus LS-Means. If you are using descriptive statistics computed from the raw data (using something like the MEANS procedure), then model estimates will not match if the sample sizes differ (i.e., the design is unbalanced).
Descriptive statistics will also differ from model estimates if the response is transformed and the model estimates are subsequently back-transformed to the original scale, or the model is a generalized linear model and model estimates are inverse-linked to the original scale.
Consequently, we need to have a certain amount of trust in the model because the model is often the best source of estimates, and that requires that we have a correctly specified model.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I see. That is good to know. So I will then report what appears as estimate as the difference value between two conditions rather than pure mean value I obtained from descriptive stat.
I learn a lot from you. Thank you very much again!
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You would want to report the estimates of the means (and SEs) which you can obtain using LSMEANS, and not just differences between means.
Another thought: Remember that GL(M)Ms report estimates on the link scale. (Estimates on the inverse-link scale can be obtained using the ILINK option.) The gamma distribution uses as log link by default, so estimates of mean (and contrasts of means) are on the log scale. Those values would clearly be different than descriptive statistics computed on the raw data.
Catch up on SAS Innovate 2026
Dive into keynotes, announcements and breakthroughs on demand.
Explore Now →ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.