- Home
- /
- Analytics
- /
- Stat Procs
- /
- Obtaining individual survival estimates
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi, I'm working on my dissertation and need to obtain individual survival estimates for everyone in my cohort. I've played around with several output commands but haven't figured out exactly what I need (i.e. baseline, xbeta, etc.). Does anyone have any advice or suggestions? One other helpful bit of information is that none of my covariates are time dependent.
Any help or advice is appreciated.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
If you want what the model estimates try looking at the outsurv= data set or basically the survival estimates.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you for your suggestion. From what I understand, the outsurv statement accompanies PROC LIFETEST, not PHREG. I need to obtain the probability of death, adjusted by covariates from my dataset. With the BASELINE statement for PHREG I have tried the SURVIVAL = _ALL_ option but my output appeared to be each covariate's mean by days of observation with survival function estimates, standard error and upper and lower confidence bounds for the survival estimate.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Then use the appropriate method for proc phreg to obtain the survival estimates.
From the doc:
output out=estimates survival;
| OUTPUT Statement |
- OUTPUT <OUT=SAS-data-set> < keyword=name ...keyword=name> </options> ;
The OUTPUT statement creates a new SAS data set containing statistics calculated for each observation. These can include the estimated linear predictor (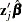 ) and its standard error, survival distribution estimates, residuals, and influence statistics. In addition, this data set includes the time variable, the explanatory variables listed in the MODEL statement, the censoring variable (if specified), and the BY, STRATA, FREQ, and ID variables (if specified).
) and its standard error, survival distribution estimates, residuals, and influence statistics. In addition, this data set includes the time variable, the explanatory variables listed in the MODEL statement, the censoring variable (if specified), and the BY, STRATA, FREQ, and ID variables (if specified).
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you for the response. I've explored the output function, too. My baseline statement is as follows:
baseline out = dataset survival = surv;
After looking at my output dataset, it appears that each covariate's values are mean values -- which isn't what I expected. Is this something I should be concerned with? With the survival function estimate by length of observation, I could go back in my original data set and merge in my estimates with observations that accumulate that length of observation, correct? I think I was thrown by expecting to see in my output dataset each patient identifier with respective 1's and 0's according to their covariate value and a survival estimate at the end. What's going on with the mean covariate values in the output dataset? Do you know anything about that?
Again, thank you for your responses.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Use the output statement NOT the baseline statement.
Here's a worked example from the SAS documentation. Your output dataset of interest is surv_data and the survival estimate is in a variable called surv_estimate. Read the doc for further information and the examples are also extremely helpful.
data Myeloma;
input Time VStatus LogBUN HGB Platelet Age LogWBC Frac
LogPBM Protein SCalc;
label Time='Survival Time'
VStatus='0=Alive 1=Dead';
datalines;
1.25 1 2.2175 9.4 1 67 3.6628 1 1.9542 12 10
1.25 1 1.9395 12.0 1 38 3.9868 1 1.9542 20 18
2.00 1 1.5185 9.8 1 81 3.8751 1 2.0000 2 15
2.00 1 1.7482 11.3 0 75 3.8062 1 1.2553 0 12
2.00 1 1.3010 5.1 0 57 3.7243 1 2.0000 3 9
3.00 1 1.5441 6.7 1 46 4.4757 0 1.9345 12 10
5.00 1 2.2355 10.1 1 50 4.9542 1 1.6628 4 9
5.00 1 1.6812 6.5 1 74 3.7324 0 1.7324 5 9
6.00 1 1.3617 9.0 1 77 3.5441 0 1.4624 1 8
6.00 1 2.1139 10.2 0 70 3.5441 1 1.3617 1 8
6.00 1 1.1139 9.7 1 60 3.5185 1 1.3979 0 10
6.00 1 1.4150 10.4 1 67 3.9294 1 1.6902 0 8
7.00 1 1.9777 9.5 1 48 3.3617 1 1.5682 5 10
7.00 1 1.0414 5.1 0 61 3.7324 1 2.0000 1 10
7.00 1 1.1761 11.4 1 53 3.7243 1 1.5185 1 13
9.00 1 1.7243 8.2 1 55 3.7993 1 1.7404 0 12
11.00 1 1.1139 14.0 1 61 3.8808 1 1.2788 0 10
11.00 1 1.2304 12.0 1 43 3.7709 1 1.1761 1 9
11.00 1 1.3010 13.2 1 65 3.7993 1 1.8195 1 10
11.00 1 1.5682 7.5 1 70 3.8865 0 1.6721 0 12
11.00 1 1.0792 9.6 1 51 3.5051 1 1.9031 0 9
13.00 1 0.7782 5.5 0 60 3.5798 1 1.3979 2 10
14.00 1 1.3979 14.6 1 66 3.7243 1 1.2553 2 10
15.00 1 1.6021 10.6 1 70 3.6902 1 1.4314 0 11
16.00 1 1.3424 9.0 1 48 3.9345 1 2.0000 0 10
16.00 1 1.3222 8.8 1 62 3.6990 1 0.6990 17 10
17.00 1 1.2304 10.0 1 53 3.8808 1 1.4472 4 9
17.00 1 1.5911 11.2 1 68 3.4314 0 1.6128 1 10
18.00 1 1.4472 7.5 1 65 3.5682 0 0.9031 7 8
19.00 1 1.0792 14.4 1 51 3.9191 1 2.0000 6 15
19.00 1 1.2553 7.5 0 60 3.7924 1 1.9294 5 9
24.00 1 1.3010 14.6 1 56 4.0899 1 0.4771 0 9
25.00 1 1.0000 12.4 1 67 3.8195 1 1.6435 0 10
26.00 1 1.2304 11.2 1 49 3.6021 1 2.0000 27 11
32.00 1 1.3222 10.6 1 46 3.6990 1 1.6335 1 9
35.00 1 1.1139 7.0 0 48 3.6532 1 1.1761 4 10
37.00 1 1.6021 11.0 1 63 3.9542 0 1.2041 7 9
41.00 1 1.0000 10.2 1 69 3.4771 1 1.4771 6 10
41.00 1 1.1461 5.0 1 70 3.5185 1 1.3424 0 9
51.00 1 1.5682 7.7 0 74 3.4150 1 1.0414 4 13
52.00 1 1.0000 10.1 1 60 3.8573 1 1.6532 4 10
54.00 1 1.2553 9.0 1 49 3.7243 1 1.6990 2 10
58.00 1 1.2041 12.1 1 42 3.6990 1 1.5798 22 10
66.00 1 1.4472 6.6 1 59 3.7853 1 1.8195 0 9
67.00 1 1.3222 12.8 1 52 3.6435 1 1.0414 1 10
88.00 1 1.1761 10.6 1 47 3.5563 0 1.7559 21 9
89.00 1 1.3222 14.0 1 63 3.6532 1 1.6232 1 9
92.00 1 1.4314 11.0 1 58 4.0755 1 1.4150 4 11
4.00 0 1.9542 10.2 1 59 4.0453 0 0.7782 12 10
4.00 0 1.9243 10.0 1 49 3.9590 0 1.6232 0 13
7.00 0 1.1139 12.4 1 48 3.7993 1 1.8573 0 10
7.00 0 1.5315 10.2 1 81 3.5911 0 1.8808 0 11
8.00 0 1.0792 9.9 1 57 3.8325 1 1.6532 0 8
12.00 0 1.1461 11.6 1 46 3.6435 0 1.1461 0 7
11.00 0 1.6128 14.0 1 60 3.7324 1 1.8451 3 9
12.00 0 1.3979 8.8 1 66 3.8388 1 1.3617 0 9
13.00 0 1.6628 4.9 0 71 3.6435 0 1.7924 0 9
16.00 0 1.1461 13.0 1 55 3.8573 0 0.9031 0 9
19.00 0 1.3222 13.0 1 59 3.7709 1 2.0000 1 10
19.00 0 1.3222 10.8 1 69 3.8808 1 1.5185 0 10
28.00 0 1.2304 7.3 1 82 3.7482 1 1.6721 0 9
41.00 0 1.7559 12.8 1 72 3.7243 1 1.4472 1 9
53.00 0 1.1139 12.0 1 66 3.6128 1 2.0000 1 11
57.00 0 1.2553 12.5 1 66 3.9685 0 1.9542 0 11
77.00 0 1.0792 14.0 1 60 3.6812 0 0.9542 0 12
;
data myeloma;
format ID 3.;
set myeloma;
id=_n_;
run;
proc phreg data=Myeloma;
model Time*VStatus(0)=LogBUN HGB Platelet Age LogWBC
Frac LogPBM Protein SCalc;
output out=surv_data survival=surv_estimate;
run;
proc print data=surv_data;
run;
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.


