- Home
- /
- Analytics
- /
- Stat Procs
- /
- Re: Interpretation of Coefficient - GLM with Gamma Link
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I have a model that requires a GLM with a log link and gamma distribution. The dependent variable is continuous and the independent variables are all dummies. The code ran for the procedure is:
PROC GENMOD DATA = TEST;
CLASS SEX_CAT BLACK_NH ASIAN_NH HISPANIC;
MODEL PRICE = SEX_CAT BLACK_NH ASIAN_NH HISPANIC / DIST = GAMMA LINK = LOG TYPE1;
WEIGHT SURVEY_WEIGHT;
RUN;
Output is as follows:
Intercept 1 6.9972 0.0513 6.8966 7.0978 18588.4 <.0001 SEX_CAT 0 1 -0.2171 0.0152 -0.2469 -0.1872 203.23 <.0001 SEX_CAT 1 0 0.0000 0.0000 0.0000 0.0000 . . BLACK_NH 0 1 0.2042 0.0226 0.1599 0.2484 81.84 <.0001 BLACK_NH 1 0 0.0000 0.0000 0.0000 0.0000 . . ASIAN_NH 0 1 0.7420 0.0347 0.6740 0.8100 457.63 <.0001 ASIAN_NH 1 0 0.0000 0.0000 0.0000 0.0000 . . HISPANIC_NH 0 1 0.7626 0.0200 0.7234 0.8018 1451.48 <.0001 HISPANIC_NH 1 0 0.0000 0.0000 0.0000 0.0000 . . Scale 1 0.3002
How would one interpret the coefficients of sex_cat or black_nh? From another SAS message board I read that:
With a log link and a continuous predictor, you are fitting the model:
ln(mu) = beta0 + beta1*X,
where mu is the expected value. Then, e raised to the left and right sides gives:
mu = exp(beta0 + beta1*X) = exp(beta0)*exp(beta1*X)
So would the interpretation for sex_cat be: exp(6.9972)(exp(-0.2171)?
Additionally, sex_cat eq 1 is female and sex_cat eq 0 equals male - so would the negative be associated with the males in comparison to females? Meaning exp(6.9972) is female, and exp(6.9972)*exp(-0.2171) is for males?
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
i would guess it's just an estimate statement with the exp option as follows
estimate 'Exp(sex_cat)' sex_cat 1 / exp cl ;
but make sure the results make sense
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Since the log function is monotonically increasing, you can interpret the parameter signs without thinking in terms of the exponentiated model. That is, the negative parameter for Sex=0 (male) indicates that being male decreases the mean response. If you need an estimate of the effect, then use the LSMEANS statement with the ILINK option: lsmeans sex / ilink;
BTW, the name of your weight variable indicates that you are attempting to analyze survey data. In general, a proper analysis can only be done using the SURVEY procedures (SURVEYMEANS, SURVEYREG, etc.) since procedures like GENMOD do not incorporate the necessary variance estimators. Unfortunately, there is no SURVEY procedure for fitting a log-linked gamma model.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
My model is similar: all independent variables are categorical. My estimates are also around 0. Could anyone give some other opinion about how to interpret the result?
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Sorry for the above reply, I thought the reply could move the topic to the top, which didn't. I'd like to ask a question about the lsmeans results interpretation:
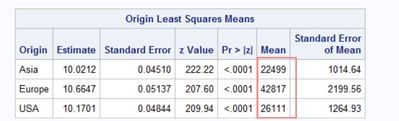
I use the sashelp.cars data to simulate an analysis. Assuming the model is the right model for the analysis. Do the lsmeans results means that compared to Asia origin, the invoice of Europe origin was 42817-22499 higher, and the invoice of USA origin was 26111-22499 higher? Thanks so much!
proc genmod data=sashelp.cars descending;
class type origin;
model invoice= type origin/dist=gamma link=log;
lsmeans origin/ilink;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I believe so, but you can easily check by adding the diff option to your LSMEANS statement.
proc genmod data=sashelp.cars descending;
class type origin;
model invoice= type origin/dist=gamma link=log;
lsmeans origin/ilink diff;
run;If there is a difference in what you calculate from the exponentiated means (and there shouldn't be in this case), it is usually due to other factors in the multiplicative model implied by a log link.
SteveDenham
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks for your reply! I have a few questions following through. Any explanation would be appreciated!
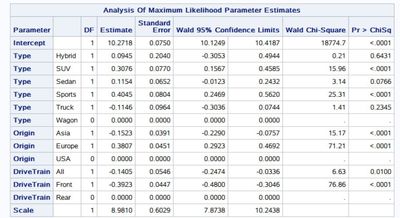
1. I calculated the exponentiated means for a few combinations(when I added another categorical variable "DriveTrain" to the model). And I'm confused about the correct way to calculate the exponentiated mean for the model. For instance, in multilinear regression, when holding other variables constant, no matter what the values other variables take(the DriveTrain could be either Front and Rear), the difference between Europe and Asia is the same. But for exponentiated results, having DriveTrain as Front and Rear shows the nonconstant difference(y1-y2≠y3-y4) as showing in the second picture. Could you please explain a little bit of how should the exponentiated means be calculated.
2. is the result of Lsmeans the same as the marginal effect? I checked there is no option to calculate its 95%CI directly, some written macro could do the work. Am I understand this right?
3. Have you used the duan's smearing estimator to calculate the mean? Any difference between the two methods(the exponentiated and the marginal effect) I mentioned above?
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
See section 4 of this note for information on properly computing the mean for any population from the parameters of the model. If you want to estimate the difference in means for a model that uses the log link (or any non-identity link), then this is not something that can be done directly in PROC GENMOD. Such a difference is a nonlinear function of the model parameters. It can be done using the NLMeans macro. See the example in the Results tab of the macro documentation. If you want to compute predicted margins or marginal effects, then use the Margins macro. See the examples in the Results tab of that macro's documentation or the links to examples provided there.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I have a question then @StatDave . The documentation for GENMOD and GLIMMIX in the LSMEANS statement clearly includes the Ilink option (and it also says that it does not report the differences on the observed scale). However, the ilink option for differences does return the ratio of the involved lsmeans under a log link, doesn't it?
I really appreciate the macros you mentioned and will be incorporating them soon.
SteveDenham
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You can use the DIFF and EXP options in LSMEANS to estimate the ratio of means in a log-linked model.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks a lot for your reply, which provides a lot of useful information! But I'm still confused when should I use Lsmean, NLSmeans macro, and the margins macro. The topic I research now is the association between comorbidity and healthcare cost. Because the independent variable of interest is a categorical variable, I guess the margins macro should work. Please correct me if I'm wrong. If anyone can provide the scenario to use all these 3 methods, that would be really appreciated!
Thanks!
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You can use the LSMEANS statement with the ILINK option to get estimates of the means for the levels of a categorical predictor, like your SEX_CAT predictor. The LSMEANS statement provides mean estimates with the other predictors fixed at their means (if continuous) or reference categories (if categorical). You can add the E option to see the coefficients used in the linear combination of model parameters that it uses to estimate each mean. The construction of LS-means is also discussed in the documentation. The margin estimates are not necessarily the same for a predictor since the other predictors do not have to be fixed. Rather, the margins are computed as the average predicted value across a copy of the data set with the predictor of interest set to each level in turn. So, the margins do not restrict the other variables to fixed values like LS-means.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
That makes sense! I'm new to this concept and your answer helps a lot! Thanks!
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand in the Innovate Hub.
Watch Now →ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.