- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi All,
Am trying to replicate a minimization from R using SAS IML. Below are sample data and IML code. The idea is to find a value for alpha that minimizes mean(gx * id)/(mean(id)**2) using the Nelder-Mead method. I should begin by saying that I’m not too familiar with SAS IML. I’ve worked my way through a number of snags but am stuck again. Some or all of my current troubles may lie with the “call nlpnms” part of the code. I’m certainly struggling with that. It may be that there are still problems occurring earlier in the code though.
data propensity;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
id = .;
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
proc iml;
use propensity;
read all var {gx};
close propensity;
start f_min(alpha) global(gx);
id = gx < alpha;
fn = mean(gx` * id)/(mean(id)**2);
return(fn);
finish f_min;
alpha = {20.3996646}; * median of gx;
optn = {0};
call nlpnms(rc, xr, "f_min", alpha, optn);
quit;
Here are the results from the calculation done in R. These suggest the value for alpha that minimizes mean(gx * id)/(mean(id)**2) should be about 24.47922.
> require(nnet)
>
> Y <- c(102, 105, 120, 130, 100, 80, 94, 108, 96)
> W <- c(1, 1, 1, 3, 2, 3, 2, 1, 2)
> X <- c(5.5, 10.6, 3.1, 8.7, 5.1, 10.2, 9.8, 4.4, 4.9)
>
> W <- relevel(as.factor(W), ref = 1)
> PF.out <- multinom(W ~ X)
# weights: 9 (4 variable)
initial value 9.887511
iter 10 value 8.236707
final value 8.235926
converged
> PF.fit <- fitted(PF.out)
>
> fn <- function (alpha)
+ {
+ gx <- apply(1/PF.fit, 1, sum)
+ id <- (gx < alpha)
+ mean(gx * id)/(mean(id)^2)
+ }
>
> optim(20.39935, fn, method = "Nelder-Mead", lower = -Inf, upper = Inf)
$par
[1] 24.47922
$value
[1] 21.02992
$counts
function gradient
6 NA
$convergence
[1] 0
$message
NULL
Thanks,
Paul
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I notice in R you are using the expression (gx * id) whereas in SAS/IML you are using (gx` * id). These are different operations. In R the '*' operator is elementwise multiplication and you use %*% for matrix multiplication. In SAS/IML, use '#' for elementwise multiplication and use '*' for matrix multiplication. For an overview of multiplication in SAS/IML, see "Ways to multiply in the SAS/IML language."
So to fix your error you can write
fn = mean(gx # id)/(mean(id)**2);
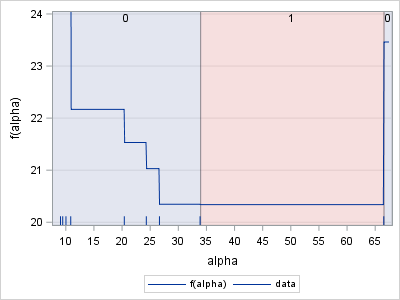
However, when I did that and re-ran your program, the NLPNMS method did not converge. There is a reason for this. Assuming that I am right about the (gx # id) expression, your objective function is a piecewise constant function. If x_i and x_{i+1} are two data values, the function is constant on (x_i, x_{i+1}]. Here is a plot of your objective function:
/* any value in (33.869, 66.526] is optimal */
alpha = do(min(gx)-1, max(gx)+1, 0.1);
y = j(1, ncol(alpha));
do i = 1 to ncol(alpha);
y[i] = f_min(alpha[i]);
end;
minVal = (y=min(y)); /* indicator variable */
create graph var {gx alpha y minVal};
append;
close;
quit;
ods graphics / width=400px height=300px;
proc sgplot data=graph;
block x=alpha block=minVal / transparency=0.8;
series x=alpha y=y / legendlabel="f(alpha)";
fringe gx / legendlabel="data";
yaxis grid min=20 max=24 label="f(alpha)";
xaxis label="alpha" values=(10 to 65 by 5) valueshint;
run;
The graph shows that the function is discontinuous and that any value in (33.869, 66.526] is a minimum. Optimization routines typically assume a continuous function with unique (local) minima and will not perform well on a discontinuous, piecewise constant function.
If you need to find the minimum of a function like this, use the fact that the function value changes at each data point and it is otherwise constant. Sort the data, and evaluate the objective function at each data point. For this example, the minimum value occurs for x=66.526. Back up to the previous data value (which is 33.869) and conclude that the optimal interval is the half-open interval (33.869, 66.526].
One more thing. You would be wise to protect yourself agains division by zero in your objective function. I would recommend that the objective function be rewritten as
start f_min(alpha) global(gx);
id = gx < alpha;
denom = mean(id)**2;
if denom > 0 then /* don't divide by 0 */
fn = mean(gx # id)/denom;
else fn = .;
return(fn);
finish f_min;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi Rick and Keshan,
Thanks very much for your replies. Rick was right in saying that I had the function specified incorrectly in IML.
Here’s the correct calculation done in IML using the initial value which is the median of gx:
data propensity;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
proc iml;
use propensity;
read all var {gx};
close propensity;
id = gx < 20.3996646;
fn = mean(gx # id)/(mean(id)**2);
print fn;
quit;
fn
21.531417
Here’s the same calculation done in R:
require(nnet)
Y <- c(102, 105, 120, 130, 100, 80, 94, 108, 96)
W <- c(1, 1, 1, 3, 2, 3, 2, 1, 2)
X <- c(5.5, 10.6, 3.1, 8.7, 5.1, 10.2, 9.8, 4.4, 4.9)
W <- relevel(as.factor(W), ref = 1)
PF.out <- multinom(W ~ X)
PF.fit <- fitted(PF.out)
fn <- function (alpha)
{
gx <- apply(1/PF.fit, 1, sum)
id <- (gx < alpha)
mean(gx * id)/(mean(id)^2)
}
fn(21.531417)
[1] 21.53139
The results seem to agree within rounding. So I think we have equivalent code. I also replicated the calculation by hand to make sure I understood what each piece of code is doing. Thanks very much Rick. This is a big step in the right direction. My apologies to Keshan if my error caused confusion. I’m not very familiar with SAS IML and so am prone to make mistakes.
So far so good. The troubling thing is how different the final answers from IML appear to be from R following minimization. As I showed in my first post, my R code and output suggest the minimum value of alpha ought to be about 24.47922 and that the value of fn corresponding to alpha ought to be about 21.02992. It turns out that the value I need is the 21.02992.
Some background information may be in order. I omitted this previously because I thought it might be unnecessary or even might constitute extraneous noise.
I’m trying to do something called generalized propensity score matching. There’s an R package called “multilevelMatching” that already does this. My company prefers a SAS-based solution. So I’m trying to replicate what this R package does using SAS. My understanding is that the two options for doing this are SAS OR and SAS IML. My company doesn’t have OR but does have IML. So I’m seeking an IML-based solution.
Part of the R code does something called “trimming”. The idea is that we should not attempt to do propensity score matching for people who have a low propensity to be in one or more treatment groups. The gx variable captures this. People with a low propensity to be in one or more treatment groups have higher scores on gx.
The R code uses the value of 21.02992 cited above to do the filtering. It multiples this value by 2 and then eliminates records with gx scores greater than this value.
So the original data based on the median of gx would look like:
data initial;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
id = gx <= 20.3996646;
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
and the data after minimization would look like:
data minimized;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
id = gx <= 21.02992 * 2;
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
Patient 3 would be trimmed because they are the only observation with a gx score > 42.05984 (21.02992 * 2). Their ID value is set to zero because they should not be included in the analysis.
Not sure why the R and SAS minimization results should be discrepant. One possbility is that here are problems in the "call nlptr" part of the code. Also think it’s possible I’m not supplying enough information. I’d certainly be happy to provide whatever information people might need.
Thanks,
Paul
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Reread my analysis again and pay particular attention to the graph of the objetive function.
If I understand the problem correctly, the value that R reports as a minimum (24.47922) is not, in fact, a minimum of the objective function, as shown in my graph. You won't be able to get an NLP function in SAS/IML to give you that value because the objective function is not continuous and so the optimization algorithm does not (and should not) converge. The Nelder-Mead simplex method (and every other NLP routine) should only be applied to continuous functions.
If I'm confused about what is going on, please explain further why you think the value 24.47922 is the value that you want.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
It is more like OR question due to some sparse variable. Post it at OR forum. @Rob is there. If I understand it rightly. You want divide gx into two group, and make Obj be min ? I think I can code some GA code for it, if you want it. I remembered a few days ago someone post a similiar question at OR forum, Check it out .
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
OK. Here is what I got .
data have;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
id = .;
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
proc sort data=have;by gx;run;
proc iml;
use have nobs nobs;
read all var {gx};
close ;
start function(x) global(gx);
xx=t(x);
call sort(xx,1,1);
if all(xx=0) then obj=99999999;
else obj=mean(gx`*xx)/((mean(xx)##2));
return (obj);
finish;
encoding=j(2,nobs,1);
encoding[1,]=0;
id=gasetup(2,nobs,123456789);
call gasetobj(id,0,"function");
call gasetsel(id,1000,1,1);
call gainit(id,10000,encoding);
niter = 1000; /*<-- Change it as large as you can to get the optimal value*/
do i = 1 to niter;
call garegen(id);
call gagetval(value, id);
end;
call gagetmem(mem, value, id, 1);
groups=t(mem);
call sort(groups,1,1);
create groups var {groups};
append;
close;
print value[l = "Min Value:"] ;
call gaend(id);
quit;
data want;
merge groups have;
run;
proc print noobs;run;
OUTPUT:
Min Value:
183.05684
GROUPS PTNO TRT PROP1 PROP2 PROP3 gx id
1 4 3 0.35394 0.34892 0.29714 9.0567 .
1 7 2 0.26402 0.29412 0.44186 9.4507 .
1 6 3 0.23172 0.26988 0.49840 10.0273 .
1 2 1 0.20073 0.24442 0.55485 10.8754 .
1 1 1 0.55445 0.38299 0.06256 20.3993 .
1 5 2 0.57192 0.37787 0.05021 24.3113 .
1 9 2 0.58017 0.37490 0.04493 26.6479 .
1 8 1 0.59957 0.36649 0.03393 33.8689 .
0 3 1 0.64347 0.34041 0.01612 66.5264 .
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I think you don't need any optimize skill, since it is sparse value.
Just enumerate all the Object Value and find that ALPHA .
data have;
input PTNO TRT PROP1 PROP2 PROP3;
gx = sum(1/prop1, 1/prop2, 1/prop3);
id = .;
cards;
3 1 0.64347 0.34041 0.01612
8 1 0.59957 0.36649 0.03393
9 2 0.58017 0.37490 0.04493
5 2 0.57192 0.37787 0.05021
1 1 0.55445 0.38299 0.06256
2 1 0.20073 0.24442 0.55485
6 3 0.23172 0.26988 0.49840
7 2 0.26402 0.29412 0.44186
4 3 0.35394 0.34892 0.29714
;
run;
proc sort data=have;by gx;run;
data want;
set have;
sum+gx;
object=sum/((_n_/9)**2);
drop sum;
run;
proc print;run;
OUPUT:
Obs PTNO TRT PROP1 PROP2 PROP3 gx id object
1 4 3 0.35394 0.34892 0.29714 9.0567 . 733.596
2 7 2 0.26402 0.29412 0.44186 9.4507 . 374.776
3 6 3 0.23172 0.26988 0.49840 10.0273 . 256.813
4 2 1 0.20073 0.24442 0.55485 10.8754 . 199.514
5 1 1 0.55445 0.38299 0.06256 20.3993 . 193.783
6 5 2 0.57192 0.37787 0.05021 24.3113 . 189.272
7 9 2 0.58017 0.37490 0.04493 26.6479 . 183.107
8 8 1 0.59957 0.36649 0.03393 33.8689 . 183.057 <--------- Minimize Value , Should be Alpha .
9 3 1 0.64347 0.34041 0.01612 66.5264 . 211.164
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Here's how to solve the problem with SAS/OR:
proc optmodel;
set OBS;
num gx {OBS};
read data propensity into OBS=[_N_] gx;
var Alpha init median(of gx[*]);
impvar Id {i in OBS} = (gx[i] < Alpha);
min Objective = (sum {i in OBS} gx[i] * Id[i] / card(OBS)) / (sum {i in OBS} Id[i] / card(OBS))^2;
solve with NLP / ms;
print Alpha Objective gx Id;
quit;
SAS Output
| Solution Summary | |
|---|---|
| Solver | Multistart NLP |
| Algorithm | Interior Point |
| Objective Function | Objective |
| Solution Status | Optimal |
| Objective Value | 20.339648389 |
| Number of Starts | 100 |
| Number of Sample Points | 400 |
| Number of Distinct Optima | 100 |
| Random Seed Used | 1337591 |
| Optimality Error | 0 |
| Infeasibility | 0 |
| Presolve Time | 0.00 |
| Solution Time | 9.51 |
| Alpha | Objective |
|---|---|
| 41.452 | 20.34 |
| [1] | gx | Id |
|---|---|---|
| 1 | 66.5264 | 0 |
| 2 | 33.8689 | 1 |
| 3 | 26.6479 | 1 |
| 4 | 24.3113 | 1 |
| 5 | 20.3993 | 1 |
| 6 | 10.8754 | 1 |
| 7 | 10.0273 | 1 |
| 8 | 9.4507 | 1 |
| 9 | 9.0567 | 1 |
Note that the optimal objective value of 20.34 agrees with @Rick_SAS's graph.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi Rob, Rick, and Keshan,
Again, thank you all for your helpful replies. I did understand Rick’s graph. I think I was just in a state of shock. I’ve been trying to process the notion that the optim function in R would produce a result that appears to be incorrect. The optim function is in the stats package, which is part of the R Core, and so I would not expect that to happen. Turns out there may be a little more to this.
What I’ve been working with is an R package called multilevelMatching. That’s not part of the R Core. It’s not even something you can download through CRAN or some other established R repository. I got the package through github. So it’s not clear to what extent it’s a work in progress, or how much checking or scrutiny it may have undergone.
I used code like the following to step through the sample call to the multilevelGPSMatch function in the multilevelMatching package.
debugonce(multilevelMatching::multilevelGPSMatch)
multilevelGPSMatch(Y, W, X, Trimming = 1, GPSM = "multinomiallogisticReg")
After working through a substantial amount of code, I was able to locate a call to the optim function from the stats package. Thing of it is though, in calling this function the multilevelMatching package turns off any warnings the optim function might provide.
Here’s what the optim function produces, including a warning.
> optim(20.39935, fn, method = "Nelder-Mead")
$par
[1] 24.47922
$value
[1] 21.02992
$counts
function gradient
6 NA
$convergence
[1] 0
$message
NULL
Warning message:
In optim(20.39935, fn, method = "Nelder-Mead") :
one-dimensional optimization by Nelder-Mead is unreliable:
use "Brent" or optimize() directly
So it’s interesting that optim is producing a value that is not in the range that Rick indicated. In its defense though, it is saying “Don’t trust this result. It’s unreliable. Try something else instead.”
The person who wrote the multilevelMatching package strikes me as an extremely intelligent and well-educated person. She has a Ph.D. in Statistics and Applied Mathematics from Iowa State University with a GPA of 4.0/4.0 and is a statistics professor at a U.S. university. So it’s interesting that she would choose to turn off the warning. We’re working with her sample data, which include only 9 records. With only these 9 records, the decision to suppress the warning doesn’t seem to turn out very well. Perhaps it would tend to work out better in more realistic situations with more data.
Thanks,
Paul
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
> Paul3 wrote:
>> "Perhaps it would tend to work out better in more realistic situations with more data."
No matter how much data you have, the objective function you've presented is discontinuous and piecewise constant. Fortunately the function changes at the data values, so there is an easy direct solution.
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand in the Innovate Hub.
Watch Now →- Ask the Expert: R and SAS Working Seamlessly Together: Unlocking PROC R in Viya for Life Sciences | 11-Jun-2026
- Ask the Expert: Can Agents Reduce Planning Friction in CPG Supply Chains? | 16-Jun-2026
- Ask the Expert: Get Meaningful Results With New Features in SAS Customer Intelligence 360 | 25-Jun-2026