- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hello Rick,
You helped me earlier to solve the problem of having negative argument inside the log function..... I was happy about to the point that I forgot that I need to carry matrix operations and this won't work due to missing values.
My question is: Can I just replace the missing values in
f = j(nrow(x), ncol(x), .); /* initialize return values to missing */
by any other small value? something like 0.01........BUT if I do so won't I be off ? Is there away that I can maybe delete the missing values well I may end up of a smaller vector than the actual size but I believe that I don't have many missing values!
Thank you,
Salah
Proc iml;
seed1=12345;
seed2=678;
seed=0;
m=10;K=1000;
a1=4; a2=5.5;
z= 1;
mu=3;lambda=2 ; *Initial values for S;
C=10;
********************** Initial values for r and S for the Bayes case *************************;
Prior_a1=J(10000,1,0); Prior_a2=J(10000,1,0);prior_g=J(10000,1,0);
Do I=1 to 10000;
Prior_a1[i]=rangam(seed,a1);
Prior_a2[i]=rangam(seed,a2);
prior_g[i]=Prior_a1[i]/(Prior_a1[i]+Prior_a2[i]);
end; *DO LOOP OF THE GAMMA;
alpha1=Prior_a1[:]; *true value for alpha1;
alpha2=Prior_a2[:]; *true value for alpha2;
gamma=alpha1/(alpha1+alpha2);
*gamma= prior_g[:];
***********************************************************************;
*************************************************************;
*** LOG likelihood
*************************************************************;
start LogLik(param) global (m,X,Y);
f=0;
alpha1=param[1];
alpha2=param[2];
c=param[3];
sum=log(1+alpha1*X+alpha2*Y) ;
f=f+m*log( alpha1*alpha2*c*(1+c) )-(c+2)*sum[+];
return(f);
finish;
**************************************************************;
/* Trap bad domain values for the Linex; return missing values when x outside domain */
start Func1(x);
f = j(nrow(x), ncol(x), .); /* initialize return values to missing */
/* trap bad domain values */
d1 = x;
domainIdx = loc( d1>0 ); /* find values inside domain */
if ncol(domainIdx)=0 then return(f); /* all inputs invalid; return missing */
t = x[domainIdx]; /* elements of t are valid */
numer = log(t);
f[domainIdx] = numer; /* evaluate f(t) */
return( f );
finish;
******************************************************************;
C_MLE=J(k,1,0); gamma_mle=J(k,1,0);S_mle=J(k,1,0);
gamma_Lin=J(k,1,0); gamma_pre=J(k,1,0);
X=J(m,1,0);Y=J(m,1,0);
Do N1=1 to K;
** Data simulation using this procedure ;
*Start Data(m,c,alpha1,alphau2,X,Y) global(seed1, seed2);
U1=J(m,1,0);U2=J(m,1,0);
do i=1 to m;
U1[i]=Uniform(seed1);
U2[i]=Uniform(seed2);
X[i]=( (1-U1[i])**(-1/c) -1 )/alpha1;*simulation is written differently;
Y[i]=(1+alpha1*X[i])/alpha2*( (1-U2[i])**(-1/(1+c)) -1 );
end;
**************************************************************************************;
*************** Optimization Step to find MLEs *********************;
************* Constrain***********************;
con={1.e-6 1.e-6 1.e-6, . . .};
opt={1};
*x0={0,0,0};
x0=alpha1||alpha2||c; *initial guess done by SAS;
tc={10000 15000};
*************** NLPQN for the Lomax function *********************;
call NLPNRA(rc, param_ret, "LogLik", x0, opt, con,tc);
alpha1_mle=param_ret[1];
alpha2_mle=param_ret[2];
c_mle[N1]=param_ret[3];
Gamma_mle[N1]=alpha1_mle/(alpha1_mle+alpha2_mle);
S_mle[N1]=(alpha1_mle+alpha2_mle);
**********************************************************************;
************************************************************************************;
************* Derivations For Lindleys method **********************;
************************************************************************************;
**L_ggg: derivative w.r.t gamma-gamma-gamma ;
**L_ggs: derivative w.r.t. gamma-gamma-S;
sum_ggg=( S_mle[N1]#(X-Y)/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##3;
L_ggg=-2*m/(1-gamma_mle[N1])**3 + 2*m/gamma_mle[N1]**3 -2*(2+c_mle[N1])* sum_ggg[+];
*********************************************************************************************************;
sum_ggs1=( 2# S_mle[N1]**2#(X-Y)##2#(gamma_mle[N1]#(X-Y)+Y ))/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) ##3 ;
sum_ggs2=( 2# S_mle[N1]#(X-Y)##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##2;
L_ggs= -(2+c_mle[N1]) *(sum_ggs1[+]-sum_ggs2[+]) ;
**********************************************************************************************************;
* L_gss was missing here ;
sum_gss1= 2# S_mle[N1]#(X-Y)#( gamma_mle[N1]#(X-Y)+Y )##2 /( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) ##3 ;
sum_gss2=2# (X-Y) #( gamma_MLE[N1]#(X-Y)+Y )/( 1+ gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2;
L_gss= -(2+c)*( sum_gss1[+] - sum_gss2[+] );
**************************************************************************************************;
**************************************************************************************************;
sum_sss=( (gamma_mle[N1]#(X-Y)+Y )/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##3;
L_sss = 2*(1+2*m)/S_mle[N1]**3 - (2+c_mle[N1])*2*sum_sss[+];
L_gsg=L_ggs; L_sgg=L_ggs;
L_ssg= L_gss ;
**************************************************************************************************;
*** Joint Prior ***;
*** P[1] is the partial derivative of p w.r.t. gamma;
*** P[2] is the partial derivative of p w.r.t. s;
P=J(2,1,0);
P[1]=(a1-1)/gamma_mle[N1]-(a2-1)/(1-gamma_mle[N1]);
p[2]=(mu-1)/S_mle[N1]-lambda;
******** Fisher Matrix *****;
sigma= J(2,2,0);
Fisher=J(2,2,0);
Sum_I11=S_mle[N1]**2#(X-Y)##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2;
Fisher[1,1]= -m/(1-gamma_mle[N1])**2 - m/gamma_mle[N1]**2 + (2+C_mle[N1])*Sum_I11[+];
Sum_I12= - S_mle[N1]#(X-Y)#( gamma_mle[N1]#(X-Y)+Y )/ ( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2
+ ( X-Y )/ ( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y );
Fisher[1,2]= -( 2+c_mle[N1] )* Sum_I12[+];
Fisher[2,1]=Fisher[1,2];
Sum_I22= ( gamma_mle[N1]#(X-Y)+Y )##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2 ;
Fisher[2,2]= - (1+2*m)/S_MLE[N1]**2 +( 2+c_mle[N1] )* Sum_I22[+];
sigma=inv(fisher);
*******************************************************************************;
***************************** Linex Loss Function ****************************;
*******************************************************************************;
U=exp(-z*gamma_mle[N1]); U_g = - z*exp(-z*gamma_mle[N1]); U_gg = z**2 *exp(-z*gamma_mle[N1]);
Exp_Lin_g = U + 0.5* U_gg* sigma[1,1] -
U_g *( sigma[1,1]*( p[1] + L_gsg* sigma[1,2] )+ sigma[1,2]*( p[2] + L_gss* sigma[1,2] ) )-
U_g/2 *( sigma[1,1]**2 *L_ggg+ sigma[1,1]*( sigma[1,2]*L_ssg+ sigma[2,2]*L_ssg)+ sigma[1,2]*L_sss*sigma[2,2] );
gamma_Lin[N1]= - Func1(Exp_Lin_g)/z;
end; * End Of Simulations;
Linex=gamma_Lin[:];
Bias_Lin=abs(gamma_Lin[:]-gamma);
MSE_Lin=(gamma_Lin-gamma_Lin[:])`*(gamma_Lin-gamma_Lin[:])/k;
print MSE_Lin bias_lin;
quit;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I'm sorry that I don't have time to carefully analyze your program. However, an initial glance causes me to wonder why Exp_Lin_g is not positive. By its name, I assume you think it is equal to (or an approximation of) EXP(t) for some t. I suggest you review that computation and check that Fisher matrix is computed correctly. The matrix should be symmetric and positive definite.
Regarding your question, the loss function is scalar, so I assume you are always calling Func1 with a scalar value and returning a scalar value. If so, the function reduces to the SafeLog function, which is Func1(x) = log(x) when x>0 and missing otherwise. At the end of the simulation, you compute the mean of gamma_Lin, so the missing values do not contribute to the mean. From what I can see, a missing value is the correct return value from Func1. However, the BEST solution is to track down and understand why Exp_Lin_g is negative.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you so much Rick for checking my problem. I believe you are right. Something is wrong with the fisher matrix.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Hi Rick,
I fixed the Fisher matrix, it is positive definite and symmetric. Unfortunately this didn't help with the values of Exp_Lin_g. I still get negative values. Exp_Lin_g must be a number between 0 and 1 as the estimate must be >0, hence - Log(Exp_Lin_g)>0
Upon checking my code carefully I was surprised to find the following:
(1) I get troubles with certain choices of the seed (negative values for Exp_Lin_g) can't tell which seeds as they keep changing!!!!
(2) I couldn't explain or understand how choosing the initials that provide the highest value for the likelihood creates errors while choices such as 1.5 and 3 which do not maximize the likelihhod function won't really work ( i.e. Exp_Lin_g<0, Exp_Lin_g depends heavily on the MLE).
Proc iml;
seed1=123;seed2=456;seed=0;
m=20; C=10;a1=2; a2=3;
********************** Initial values for alpha1 and alpha2 for the Bayes case *************************;
Prior_a1=J(10000,1,0); Prior_a2=J(10000,1,0);prior_g=J(10000,1,0);
Do I=1 to 10000;
Prior_a1[i]=rangam(seed,a1); Prior_a2[i]=rangam(seed,a2); prior_g[i]=Prior_a1[i]/(Prior_a1[i]+Prior_a2[i]);
end;
alpha1=Prior_a1[:]; alpha2=Prior_a2[:]; gamma=prior_g[:];
*********************************************************************;
/* write the log-likelihood function for bivariate Lomax dist */
***********************************************************************;
start LogLik(param) global (m,X,Y);
f=0; alpha1=param[1]; alpha2=param[2]; c=param[3];
sum=log(1+alpha1*X+alpha2*Y) ;
f=f+m*log( alpha1*alpha2*c*(1+c) )-(c+2)*sum[+];
return(f);
finish;
*****************************************************************************;
** Simulating data from bivariate Lomax with parameters alpha1, alpha2 and C ;
*****************************************************************************;
X=J(m,1,0);Y=J(m,1,0);
U1=J(m,1,0);U2=J(m,1,0);
do i=1 to m;
U1[i]=Uniform(seed1);
U2[i]=Uniform(seed2);
X[i]=( (1-U1[i])**(-1/c) -1 )/alpha1; ** Simulating X from f(x);
Y[i]=(1+alpha1*X[i])/alpha2*( (1-U2[i])**(-1/(1+c)) -1 ); ** Simulating y from f(Y|x);
end;
/* optional: test the module */
params = { 1.5 3 10, 0.9 0.9 10, 0.9 .1 10, 0.1 0.9 10, 0.7 0.7 10, 0.7 1 10,4 6 10,5 4 10, 5.5 4 10,
1 0.7 10, 1.5 5 10, 1.5 5.5 10, 1.5 6 10, 5 1.5 10, 5.5 1.5 10, 6 1.5 10,5 5 10,5 5.5 10,
2 2 10, 2 5 10, 2 5.5 10, 2 6 10, 5 2 10, 5.5 2 10, 6 2 10, 6 4 10, 5 6 10,
3 5 10, 3 5.5 10, 3 6 10, 5 3 10, 5.5 3 10, 6 3 10,4 5 10,4 5.5 10,5.5 5 10,6 5 10};
LogLik = j(nrow(params),1);
do i = 1 to nrow(params);
p = params[i,];
LogLik[i] = LogLik(p);
end;
results = LogLik || Params;
call sort(results, 1, 1); /* decreasing by loglik */
print (results[1:30, ])[c={"LogLik" "alpha1" "alpha2" "C"} L="results"];
call scatter(1:nrow(results), results[,1]);
quit;
(3) Increasing the stimulation size results on increasing the chances of getting awful values for Exp_Lin_g...I mean getting large negative/positive values such as -2.12 or 3.7!!!!!
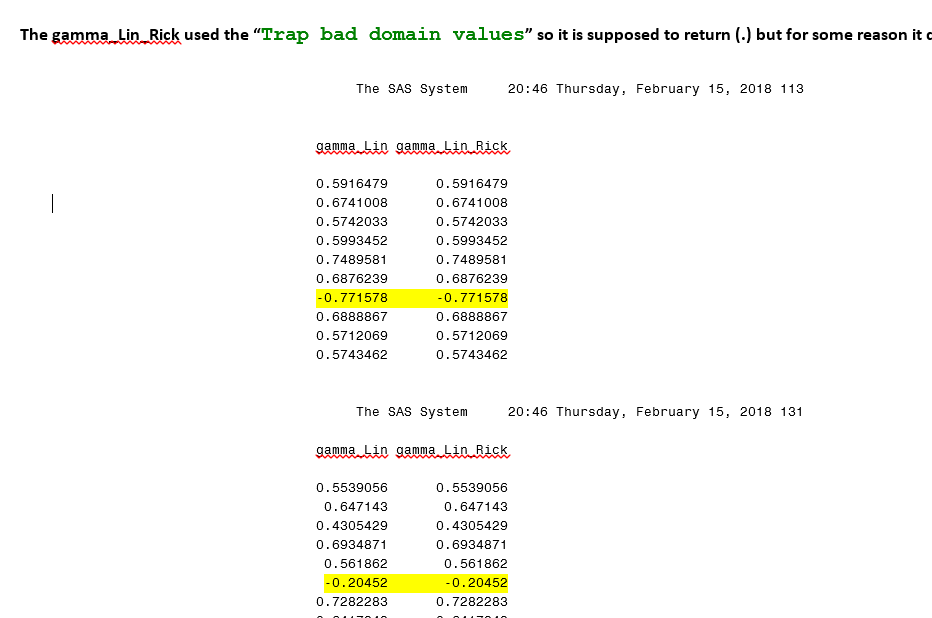
(4) Replacing negative values with missing values is not working. I called it "gamma_lin_Rick and as you see in the pic below it is negative and it supposed to be "."!!!!
I am attaching my code below. Thank you Rick.
Proc iml;
*seed1=2; *seed2=22; *These choices are working sometimes;
*seed1=123; *seed2=0; seed1=123; seed2=456;
m=10;K=1000;
a1=1.5;a2=2;
a1=1.5; a2=3; /*this choice caused mistakes*/
C=10;z= 1;mu=3;lambda=2 ; *Initial values for S;
* Initial values for r and S for the Bayes case **;
Prior_a1=J(10000,1,0); Prior_a2=J(10000,1,0);prior_g=J(10000,1,0);
Do I=1 to 10000;
Prior_a1[i]=RAND('GAMMA',a1);
Prior_a2[i]=RAND('GAMMA',a2);
prior_g[i]= RAND('BETA',a1,a2);
end; *DO LOOP OF THE GAMMA;
alpha1=Prior_a1[:]; *true value for alpha1;
alpha2=Prior_a2[:]; *true value for alpha2;
gamma= prior_g[:];
***********************************************************************;
*** Log likelihood function
************************************************************************;
start LogLik(param) global (m,X,Y);
f=0; alpha1=param[1]; alpha2=param[2]; c=param[3];
sum=log(1+alpha1*X+alpha2*Y) ;
f=f+m*log( alpha1*alpha2*c*(1+c) )-(c+2)*sum[+];
return(f);
finish;
**************************************************************;
/* Trap bad domain values; return missing values when x outside domain */
start Func1(x);
f = j(nrow(x), ncol(x), .); /* initialize return values to missing */
d1 = x;
domainIdx = loc( d1>0 ); /* find values inside domain */
if ncol(domainIdx)=0 then return(f); /* all inputs invalid; return missing */
t = x[domainIdx]; /* elements of t are valid */
numer = log(t);
f[domainIdx] = numer; /* evaluate f(t) */
return( f );
finish;
/* The part below is for simulating data from Bivariate Lomax*/
C_MLE=J(k,1,0); gamma_mle=J(k,1,0);S_mle=J(k,1,0);
gamma_Lin=J(k,1,0); gamma_Lin_Rick=J(k,1,0);Exp_Lin_g=J(k,1,0);
X=J(m,1,0);Y=J(m,1,0);
Do N1=1 to K;
U1=J(m,1,0);U2=J(m,1,0);
do i=1 to m;
U1[i]=Uniform(seed1);
U2[i]=Uniform(seed2);
/* the method below is a better way for simualtion,I wanted to control the seed so I can figure out what went wrong!!!!!
/* But I failed!!!!!!
*U1[i]=RAND('UNIFORM');
*U2[i]=RAND('UNIFORM');
*/
X[i]=( (1-U1[i])**(-1/c) -1 )/alpha1;
Y[i]=(1+alpha1*X[i])/alpha2*( (1-U2[i])**(-1/(1+c)) -1 );
end;
/* End of the simulation */
*************** Optimization Step to find MLEs *********************;
************* Constrain***********************;
con={1.e-6 1.e-6 1.e-6, . . .};
opt={1};
x0=alpha1||alpha2||c; *initial guess done by SAS;
tc={10000 15000};
*************** NLPQN for the Lomax function *********************;
call NLPNRA(rc, param_ret, "LogLik", x0, opt, con,tc);
alpha1_mle=param_ret[1];
alpha2_mle=param_ret[2];
c_mle[N1]=param_ret[3];
Gamma_mle[N1]=alpha1_mle/(alpha1_mle+alpha2_mle);
S_mle[N1]=(alpha1_mle+alpha2_mle);
************************************************************************************;
************* Derivations For Lindleys method **********************;
************************************************************************************;
**L_ggg: derivative w.r.t gamma-gamma-gamma ;
**L_ggs: derivative w.r.t. gamma-gamma-S;
sum_ggg=( S_mle[N1]#(X-Y)/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##3;
L_ggg=-2*m/(1-gamma_mle[N1])**3 + 2*m/gamma_mle[N1]**3 -2*(2+c_mle[N1])* sum_ggg[+];
*************************************************************************************;
sum_ggs1=( 2# S_mle[N1]**2#(X-Y)##2#(gamma_mle[N1]#(X-Y)+Y ))/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) ##3 ;
sum_ggs2=( 2# S_mle[N1]#(X-Y)##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##2;
L_ggs= -(2+c_mle[N1]) *(sum_ggs1[+]-sum_ggs2[+]) ;
**************************************************************************;
sum_gss1= 2# S_mle[N1]#(X-Y)#( gamma_mle[N1]#(X-Y)+Y )##2 /( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) ##3 ;
sum_gss2=2# (X-Y) #( gamma_MLE[N1]#(X-Y)+Y )/( 1+ gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2;
L_gss= -(2+c)*( sum_gss1[+] - sum_gss2[+] );
*********************************************************************;
********************************************************************;
sum_sss=( (gamma_mle[N1]#(X-Y)+Y )/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y ) )##3;
L_sss = 2*(1+2*m)/S_mle[N1]**3 - (2+c_mle[N1])*2*sum_sss[+];
L_gsg=L_ggs; L_sgg=L_ggs;
L_ssg= L_gss ;
***********************************************************************************;
*** Joint Prior ***;
P=J(2,1,0);
P[1]=(a1-1)/gamma_mle[N1]-(a2-1)/(1-gamma_mle[N1]);
p[2]=(mu-1)/S_mle[N1]-lambda;
******** Fisher Matrix *****;
sigma= J(2,2,0);
Fisher=J(2,2,0);
Sum_I11=S_mle[N1]**2#(X-Y)##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2;
Fisher[1,1]= -(-m/(1-gamma_mle[N1])**2 - m/gamma_mle[N1]**2 + (2+C_mle[N1])*Sum_I11[+]);
Sum_I12= - S_mle[N1]#(X-Y)#( gamma_mle[N1]#(X-Y)+Y )/ ( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2
+ ( X-Y )/ ( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y );
Fisher[1,2]= ( 2+c_mle[N1] )* Sum_I12[+];
Fisher[2,1]=Fisher[1,2];
Sum_I22= ( gamma_mle[N1]#(X-Y)+Y )##2/( 1+gamma_MLE[N1]#S_MLE[N1]#(X-Y)+S_MLE[N1]#Y )##2 ;
Fisher[2,2]= (2*m)/S_MLE[N1]**2 -( 2+c_mle[N1] )* Sum_I22[+];
sigma=inv(fisher);
*print (eigval(sigma)); /*this is to check is sigma is positive definite*/
*******************************************************************************;
***************************** Linex Loss Function ****************************;
*******************************************************************************;
U=exp(-z*gamma_mle[N1]); U_g = - z*exp(-z*gamma_mle[N1]); U_gg = z**2 *exp(-z*gamma_mle[N1]);
x1=0.5* U_gg* sigma[1,1];
x2=sigma[1,1]*( p[1] + L_gsg* sigma[1,2] );
x3=sigma[1,2]*( p[2] + L_gss* sigma[1,2] );
x4=sigma[1,1]**2 *L_ggg;
x5=sigma[1,1]*( sigma[1,2]*L_ggs+ sigma[2,2]*L_ssg);
x6=sigma[1,2]*L_sss*sigma[2,2];
Exp_Lin_g[N1] = U + x1 + U_g *( x2 + x3 )+ 0.5 *U_g *( x4+ x5+ x6 );
gamma_Lin_Rick[N1]= -Func1(Exp_Lin_g[N1])/z;
gamma_Lin[N1]= -Log(Exp_Lin_g[N1])/z;
end; * End Of Simulations;
Estimate_Linex=gamma_Lin[:];
MSE=(gamma_Lin-gamma_Lin[:])`*(gamma_Lin-gamma_Lin[:])/k;
print Estimate_Linex " " MSE;
print gamma_Lin gamma_Lin_Rick ;
quit;
April 27 – 30 | Gaylord Texan | Grapevine, Texas
Registration is open
Walk in ready to learn. Walk out ready to deliver. This is the data and AI conference you can't afford to miss.
Register now and lock in 2025 pricing—just $495!
- Ask the Expert: Wie kann ich Visual Studio Code als SAS Entwicklungsumgebung nutzen? | 11-Dec-2025
- DCSUG Online Winter Webinar | 11-Dec-2025
- Ask the Expert: Marketing Decisions Excellence: Turning Insights Into Business Impact | 16-Dec-2025
- SAS Bowl LVII, SAS Data Maker and Synthetic Data | 17-Dec-2025
- SAS Innovate 2026 | Grapevine, Texas | 27-Apr-2026