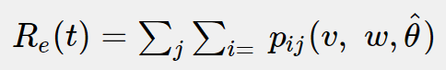- Home
- /
- SAS Communities Library
- /
- SAS Helps You Understand Disease Spread: Part 2 Biostat Concepts in th...
- RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
SAS Helps You Understand Disease Spread: Part 2 Biostat Concepts in the Time of Coronavirus
- Article History
- RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
In this time of the Covid-19 pandemic, understanding how disease spreads can save lives. As for many of us, this virus has touched me in a personal way.
Flashback to February of 2020. My good friend Cristina’s parents were getting on in years, but still managing to live on their own here in my home town. Her 89-year-old mother has dementia and so her charismatic 92-year old father was taking care of things. They needed some extra help so they hired a woman named Sofia (not her real name) to come in for 2 hours a day to help with meals and such. Unfortunately, Sofia came down with Covid-19, ending up on a ventilator for weeks in the hospital. My friend’s parents both contracted the disease. Thanks to the great care of Cristina and her sister, Cristina’s mother survived. After many weeks on a ventilator, Sofia also survived. Unfortunately, my friend’s father went to the hospital, and never came home.
I have four other friends (that I know of) who have come down with Covid-19. Three have recovered. One of them, close to my own age, also died. In Maryland (my state of about 6 million) we lost 11 more Marylanders yesterday from Covid-19 (September 23). While terrible, 11 deaths is much better than our peak of 68 deaths in one day on April 29.
Understanding the spread of an infectious disease, as explained in this article, can help us to reduce that spread and reduce the number of illnesses and deaths. Check out articles on other biostats concepts in this series: Sensitivity & Specificity in Disease Testing: Part 1 Statistical Concepts in the Time of Coronavirus...and Network Analysis in SAS Visual Analytics: Part 3 of Biostats in the Time of Coronavirus.
Mobile users: To view the images, select the "Full" version at the bottom of the page.
Background
Infectious diseases have affected humans throughout recorded history. They are caused by pathogens, including viruses or bacteria. A virus is a nucleic acid strand (RNA or DNA) protected by a protein shell, and sometimes enveloped by an external membrane similar to the plasma membrane of a host cell. Outside of their hosts, viruses break down quickly and cannot survive for long. Densely settled human populations provide prime conditions for spreading diseases, allowing bacteria and viruses to spread between humans, and also between animals and humans. Some notable diseases that have dramatically affected human history include:
- Plague
- Influenza
- Measles
- Smallpox
The plague is famous in Europe for causing the epidemic dubbed the “Great Pestilence” or “Black Death.” It killed roughly a third of the people in Europe in the 14th Century. Plague is caused by the bacterium Yersinia pestis, which travels from person to person through the air or from infected flea or rat bites. Gross! In October 1347, twelve rat-infested ships from the Black Sea docked at a Sicilian port. Many of the sailors on board were dead or covered in boils. Ultimately the spread of the plague was slowed by forcing sailors to remain on their ships for 40 days. Our modern word quarantine originates from the term quarantena, which means forty days. There are still between one and three thousand cases of plague a year to this day!
The influenza pandemic of 1918, frequently dubbed the “Spanish Flu,” killed an estimated 20 to 50 million people world wide. For comparison, 17 million people died in the First World War. The first known case of the “Spanish” Flu was actually reported at a military base in Kansas, US on March 11, 1918. Around 675,000 Americans died; at the time the US population was around 100 million. In the US, tens of thousands still die annually of influenza, perhaps as many as 62,000 in the 2019-2020 flu season.
The smallpox virus killed 3 out of every 10 people in Eurasia for centuries. When European explorers and conquistadors seeking fortunes for European royalty came to the Americas in the 15th century, they brought the virus with them. The Native Americans were 100% susceptible, having no antibodies to the virus in their population. Tens of millions of people died and 90 to 95 percent of the indigenous population was wiped out. Although a smallpox vaccine was invented in the late 1700s, there were still around 50 million smallpox deaths a year in the 1950s. But good news! Smallpox was officially eradicated worldwide by 1980.
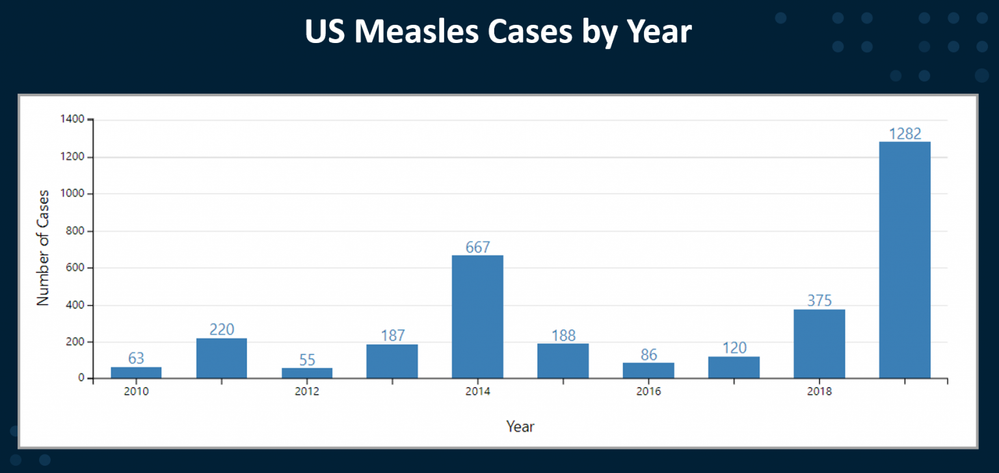
Measles is a highly contagious disease caused by a virus. Worldwide, measles kills more than 140,000 people a year; the majority of these deaths are in countries with low per capita incomes and weak health infrastructures. The measles virus hitches a ride in droplets from the nose and mouth of the infector to the infectee. Symptoms commonly appear 10-12 days later. Measles was declared eliminated from the US in 2000 but recent years have seen a resurgence. The measles vaccine is considered highly effective; one dose is 93% effective; two doses are 97% effective. In my state of Maryland, a measles vaccine is required for children to enroll in school.
Some notable infectious diseases that have caused concerning epidemics more recently include:
- Ebola
- Coronaviruses
- SARS
- MERS
- Covid-19
Terms
The difference between outbreaks, epidemics and pandemics is based on size and geographic distribution.
- outbreak - occurs in a limited area where a number of disease cases is larger than what you would expect for a given population in a given time frame
- epidemic - when an outbreak spreads over a large number of people in a wide geographic area in a short period of time
- pandemic - a global epidemic, many parts of the world are affected at once
Factors that affect the spread of disease include the reproduction number R0 and the serial interval.
- reproduction number R0 - how many people an infected person will transmit the disease to; more precisely, it is the number of secondary infections produced by an infected individual in a population that is totally susceptible
- effective reproduction number Re - the average number of secondary cases produced by a primary case; it may be calculated as in the following formula:
- serial interval - time from onset of symptoms in the infector to the onset of symptoms in the infectee; serial interval differs from incubation period because incubation period does not account for time of symptom onset in the infector
- incidence - the number of new cases per time unit (e.g., per day, per year) divided by the total population number; incidence tells us how quickly people are becoming affected
- prevalence - how much of a population is affected at a given time (per 100,000 people)
Some infectious diseases may be spread by non-human animals:
- zoonotic disease - infectious disease caused by a pathogen (such as a bacterium, virus, parasite or fungus) that has jumped from an animal (such as a cow, pig, bat, or rat) to a human; zoonotic commonly refer to diseases that can jump from vertebrate animals
- vector-borne disease - include diseases that can be transmitted by mosquitoes, ticks, and fleas, such as Lyme Disease and Zika virus
The disease Covid-19 is caused by a type of coronavirus. The virus itself is named the novel SARS-CoV-2. There is still much to learn about this virus. More information is known about two other coronaviruses is known because they have been studied longer. For example, we know that MERS-CoV is widespread in dromedary camels. Neither SARS nor MERS became a worldwide pandemic.
Spread
Several things may increase concern about us about a disease:
- How serious is the disease?
- How easily does it spread?
- How much of the population is susceptible?
Case Fatality Rate (CFR) is how many people die out of the number of people who have confirmed cases of the disease. Note that CFR is not the same as the mortality rate, which is the number of deaths per 100,000 persons in the total population. CFR can vary by a number of factors such as age group and the availability of quality health care.
Case fatality rates for Covid-19 have been reported from 1.6% to 10.5%.
How easily it spreads
If the transmission risk of the disease is 100% and each infectious person meets two other susceptible people before the infector recovers, the disease will soon begin to spread very quickly. Assuming that recovery takes one day, this situation will result in the number of sick people doubling each day. In this situation:
y = 2t−1
where:
y = the total number of people infected, and
t = the time, in days, which has elapsed since the initial outbreak.
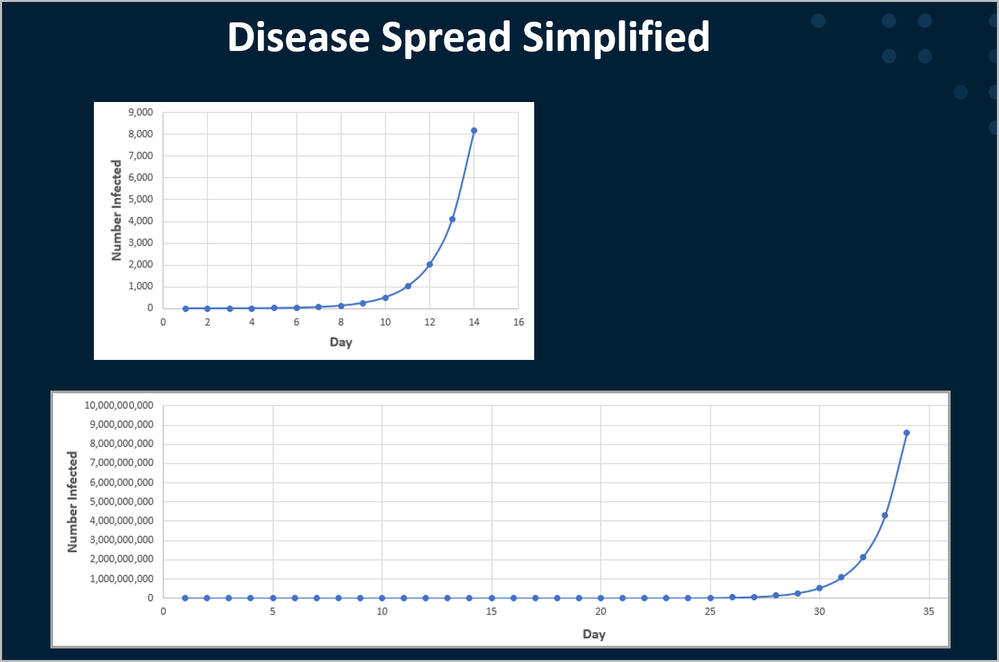
In this highly simplified example, we would see graphically:
Thus we would see 8,000 infected people in 8 days, and over 8 billion (the whole world) infected in less than 35 day.
But! This is not how networks work. For example, Person A may infect Person B. Then Person A and Person B may come into contact with Person C. But that is only one infection in Person C, not 2 infections. These complications are why using network analysis tools can help you!
Reproduction Number R0
The reproduction number (R0) is the average number of people an infected person will transmit the disease to in a fully susceptible population. For example, if the value of R0 for measles is 14, then each case of measles would produce 14 new secondary cases. This would spread through the population much faster than a disease where the value of R0 is only equal to 2.
If R0 > 1 then the infection will spread and, unchecked, will lead to an epidemic. On the other hand, if R0 < 1, the infection will likely die out. The higher the R0 the more quickly the disease will spread, and the more difficult it will be to contain an outbreak and prevent an epidemic.
R0 is affected by:
- The rate of contact between individuals in the host population
- The probability of the infection being transmitted during contact
- The duration of infectiousness
Here’s the good news. Interventions can reduce the reproduction number by reducing contact among individuals and reducing the probability of infection being transmitted. Social distancing, stay-at-home and safer-at-home orders, and mask wearing have been recommended in most countries to help reduce R0.
Now that we are familiar with these terms, let’s compare some example estimates of serial interval, R0, and CFR from the literature for different diseases:
Herd Immunity
Herd immunity has to do with increased resistance of a population to a disease outbreak or epidemic due to a high number if individuals in the population having immunity, which decreases the likelihood of an infected individual coming into contact with a susceptible individual.
Different folks define herd immunity differently. Some define it as partial resistance, reflected in reductions in frequency of disease due to fewer infectors and fewer susceptible individuals. Some define it as total resistance, such as a percentage of immune individuals in a population above which a disease will die out.
If the proportion of immune individuals is high enough that the number of susceptible individuals is below the epidemic threshold, then incidence will decrease. This herd immunity threshold can be calculated as follows.
Graphed, this would look like:
Using this formula, and the R0s in the table up above, we can estimate Herd Immunity Thresholds.
If we assume a CFR of only 1.6%, to achieve 50% herd immunity (without a vaccine), that would translate into over 2 million US deaths (2,648,000) or over 62 million world deaths. To achieve 87% herd immunity, that would translate into 4.6 million US deaths or over 108 million deaths worldwide.
Network Analysis Can Inform Strategies to Reduce Spread
We can use network analysis in a variety of ways to help determine:
- What areas and what individuals are likely to be hit next
- Where and when limited resources should be applied
- Where facilities or individuals might be superspreaders; these locations and individuals might best be targeted with information campaigns, elevated testing opportunities, et cetera
- What areas can be protected
- How can we lower deaths and hospitalizations
SAS provides a number of network analysis tools to help you!
These include:
- SAS Visual Analytics
- SAS VDMML
- SAS Visual Investigator
- SAS Viya with Python
1. SAS Visual Analytics
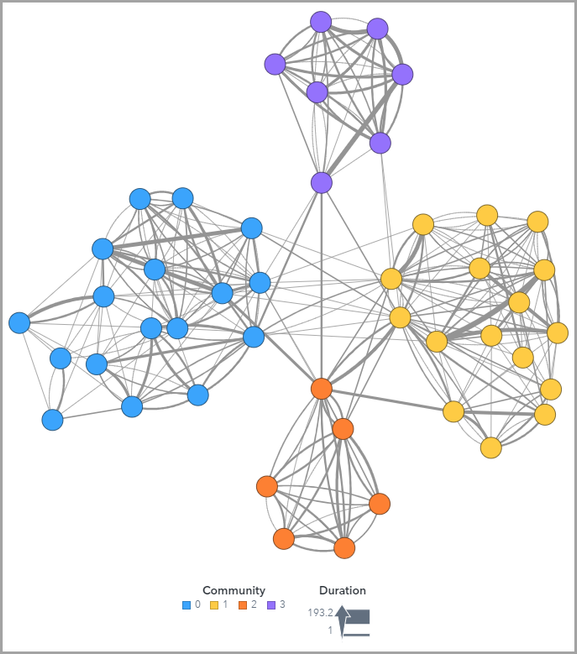
The SAS Visual Analytics Network Analysis object uses node color and size and link color and size to let you visualize network metrics. It can identify communities (as shown below) where individuals are more linked to each other, and other network metrics such as reach centrality, closeness centrality, stress centrality and betweenness centrality. See my next article to learn how to use the SAS Visual Analytics Network Analysis object. Sneak peek below.
The Visual Analytics Network Analysis object uses the Hypergroup Action Set, which includes two actions.
| Action | Description |
| Hypergroup | Find hypergroups, graph layouts, colors, communities, centralities, and create structural and nBody graphs. Find shortest paths. |
| thePlotThickens | Determine which points to display in plots when the data has a great many points. |
For more information, see the documentation.
2. SAS Visual Data Mining and Machine Learning (VDMML)
VDMML allows you to use PROC NETWORK, which is based on the Network Action Set. The actions with that set are listed below.
In the VDMML 8.5 release, enhancements reduced the computation time for some of the algorithms. For additional details, see the documentation.
Quite a few examples are available to you.
For more information, see the documentation.
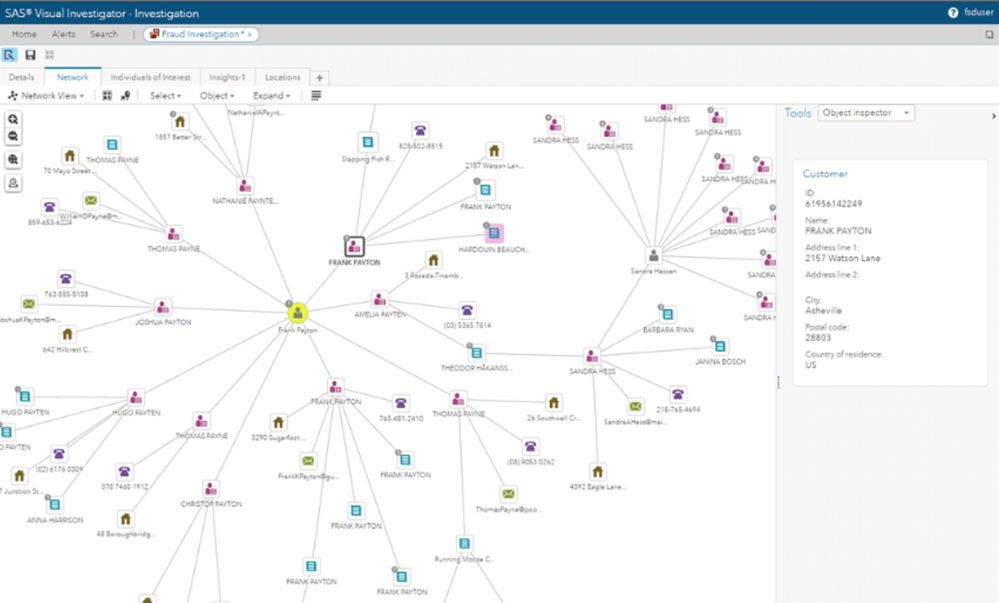
3. SAS Visual Investigator
SAS Visual Investigator provides highly customizable social network analysis tools.
4. SAS Viya with Python
The application developer or data scientist with Python programming experience can use Python with SAS Viya to analyze a social network.
In summary, understanding the spread of disease can help us save lives and reduce adverse effects. SAS has many social network analysis tools to help us understand the spread of disease. See my next article to get started by using the SAS Visual Analytics Network Analysis object.
Extra Tips from Beth
You’re welcome. 😊
For More Information:
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Great article, thank you! Could you please add examples of simulations using R0 and incubation time in SAS? Best wishes!
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Thanks @BethEbersole for sharing this helpful article and I'm sorry to hear how this virus has tragically affected you and your friends.
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →SAS AI and Machine Learning Courses
The rapid growth of AI technologies is driving an AI skills gap and demand for AI talent. Ready to grow your AI literacy? SAS offers free ways to get started for beginners, business leaders, and analytics professionals of all skill levels. Your future self will thank you.
- Find more articles tagged with:
- COVID-19 biostats