- Home
- /
- Analytics
- /
- SAS Data Science
- /
- Re: SAS Miner: Export Leaf Table (without manually saving) from a node...
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I have a large project in which I generated 400+ decision trees. I need to export the SAS Node Leaf dataset automatically from a node. I do not want to have to click "Save as" 400+ times. Is there a way I can export it as part of the model? Placing the control point at the end of the all the paths has been a great tool to generate all the trees "at once". Can I insert a node to export the leaf datasets before the control point?
Also, is there a way to do this with the output which contain the fit statistics?
Thanks.
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I believe the correct names of the data sets are actually TreeZ_outseq, TreeZ_outnodes, and TreeZ_outstats (assuming that is what corresponds to statsbynode). You can see all the data sets from the Decision Tree node by opening the Explorer window inside EM, click the checkbox for Show Project Data, then navigate to your workspace folder (Emws15 e.g.). All the data sets with the TreeZ prefix are the ones created by the Decision Tree node.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The EM Decision Tree node saves the tree model in a data set that PROC ARBOR understands. PROC ARBOR can output information about all the nodes in the tree. It has no output just for the leaves.
In an EM code node,
PROC ARBOR INMODEL= &em_lib..treeZ_emtree;
MACRO NLEAVES=nleaves;
SAVE SEQUENCE=sequence NODESTATS=nodes STATSBYNODE=statsbynode;
QUIT;
inputs the tree &em_lib..treeZ_emtree, where TreeZ is the node ID for the Decision Tree node, and outputs
a macro variable, NLEAVES, equal to the number of leaves in the tree, and three data sets:
sequence: each row has fit statistics for a subtree with a specific number of leaves,
nodes: each row contains statistics for a node (including leaves) in the current subtree (with &NLEAVES leaves)
statsbynode: node statistics re-arranged to allow for more statistics.
(I am not 100% sure about the macro name, &em_lib..treeZ_emtree.)
Is this what you need to know?
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
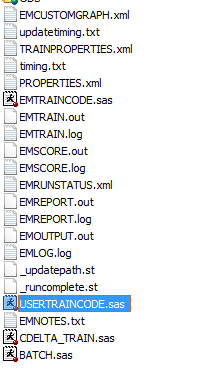
This is exactly what I need! However, when I run the below code in the SAS code node, I am not seeing the three data sets you mentioned. I only see the file names below. Any thoughts? Thanks.
.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I would first look at the log. I do not know enough about the EM interface to find the log. If no one else responds, try contacting tech support.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The logs have an error when looking for the EMTREE data.
Maybe I don't understand your notation for "TREEZ", but I tried substituting the node id (Tree38) for Z
ERROR: File WORK.EMWS15TREE38_EMTREE.DATA does not exist
and when I leave in TREEZ
ERROR: File WORK.EMWS15TREEZ_EMTREE.DATA does not exist
Either way, I don't see any *EMTREE* data in my directories.
Any idea?
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
You're helping me more than I am helping you now. The EM macro, directory, and file names are a mystery to me. I'm sorry I can't help more.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Make sure you use 2 periods before tree38:
inmodel=&em_lib..tree38_emtree
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Yes, that was my issue! However, I still don't see the files for "node", "sequence", or "nodes" my EM_LIB directory. That should be the names of the datasets. Correct?
Thank you!
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The log should report the name of the dataset and the number of obsevations shortly after it reports the PROC ARBOR and SAVE statement. Remember to replace "sequence" by a libname.memname of your choice. For example, EM_LIB.Sequence.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thanks. I tried "&em_lib..", "&em_lib.", and "em_lib." as repressented by the below in bold:
PROC ARBOR INMODEL= &em_lib..Tree38_emtree;
MACRO NLEAVES= &em_lib..nleaves;
SAVE SEQUENCE= &em_lib..sequence NODESTATS= &em_lib..nodes STATSBYNODE= &em_lib..statsbynode;
QUIT;
the log says (and no location):
14896 %let syscc = 0;
14897 %inc "/xxxx/Projects/xxxx Data Mining 2015/Predictive Models/Workspaces/EMWS15/EMCODE/EMTRAINCODE.sas";
NOTE: %INCLUDE (level 1) file /xxxx/Projects/xxxx Data Mining 2015/Predictive Models/Workspaces/EMWS15/EMCODE/EMTRAINCODE.sas is file /xxxxx/Projects/xxxxx Data Mining 2015/Predictive Models/Workspaces/EMWS15/EMCODE/EMTRAINCODE.sas.
14898 +PROC ARBOR INMODEL= &em_lib..Tree38_emtree;
NOTE: 1654978 kilobytes of physical memory.
NOTE: The subtree sequence contains 5 subtrees. The largest has 9 nodes and 5 leaves.
14899 +MACRO NLEAVES= &em_lib..nleaves;
14900 +SAVE SEQUENCE= &em_lib..sequence NODESTATS= &em_lib..nodes STATSBYNODE= &em_lib..statsbynode;
14901 +QUIT;
NOTE: %INCLUDE (level 1) ending.
14902 *------------------------------------------------------------*;
14903 * Close any missing semi colons;
14904 *------------------------------------------------------------*;
14905 ;
14906 ;
14907 ;
14908 ;
14909 quit;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I received an email asking if I have a solution. I still do not. I am trying many options including the ones I highlighted in my previous post.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I believe the correct names of the data sets are actually TreeZ_outseq, TreeZ_outnodes, and TreeZ_outstats (assuming that is what corresponds to statsbynode). You can see all the data sets from the Decision Tree node by opening the Explorer window inside EM, click the checkbox for Show Project Data, then navigate to your workspace folder (Emws15 e.g.). All the data sets with the TreeZ prefix are the ones created by the Decision Tree node.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Thank you!! Exactly what I was looking for. I can't believe how hard I made this.
Available on demand!
Missed SAS Innovate Las Vegas? Watch all the action for free! View the keynotes, general sessions and 22 breakouts on demand.
Use this tutorial as a handy guide to weigh the pros and cons of these commonly used machine learning algorithms.
Find more tutorials on the SAS Users YouTube channel.