- Home
- /
- Analytics
- /
- Stat Procs
- /
- Bayesian TTest by Proc MCMC
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
This t-Test is taken from Statistical Business Analysis Using SAS
libname sasba 'c:\sasba\ames';
data ames;
set sasba.ames300;
run;
proc format;
value yesno 0=No 1=Yes;
run;
proc ttest data =ames plots (only)=qq alpha=.05 h0=0;
class bonus;
var gr_liv_area;
format bonus yesno.;
run;
Let's imagine we want to repeat the test using Bayes, in BUGS we could approach/reproduce! that result with this model.
model {
for (i in 1:2) {
mu[i] ~ dnorm(0.0,1.0E-6)
tau[i] ~ dgamma(0.001,0.001)
}
for (i in 1:n) {
area[i] ~ dnorm(mu[bonus[i]+1],tau[bonus[i]+1])
}
x1 ~ dnorm(mu[1],tau[1])
x2 ~ dnorm(mu[2],tau[2])
dx<- x1-x2
dmu<-mu[1]-mu[2]
}
list(
area = c(864, 1829, 1328, 1063, 2207, 972, 912, 1978, 1801, 2018, 882, 1370, 1350, 2402, 1600, 2020, 1629, 2452, 2490, 1114, 864, 1740, 1728, 1313, 1154, 1113, 1567, 1392, 1144, 1478, 1297, 1062, 1121, 1092, 1513, 1368, 1560, 1680, 2036, 1641, 874, 1782, 2464, 1574, 1460, 1175, 1740, 925, 1136, 1092, 1100, 1196, 1044, 825, 1117, 882, 1211, 1524, 1094, 1418, 924, 1445, 1091, 1279, 923, 816, 914, 872, 2633, 1636, 988, 1093, 1214, 1150, 912, 1442, 1721, 922, 948, 952, 1242, 897, 955, 1299, 998, 1342, 1442, 1500, 907, 1214, 768, 1839, 1680, 1696, 1478, 2279, 1240, 1040, 1293, 1868, 1780, 2156, 1334, 1251, 1216, 1124, 884, 1045, 1073, 1159, 1458, 1124, 1654, 1339, 720, 1068, 1296, 1022, 952, 875, 520, 838, 672, 816, 778, 968, 960, 1047, 694, 1103, 693, 641, 865, 884, 1108, 1616, 1536, 1962, 1154, 1668, 1374, 1461, 1328, 1324, 1412, 1112, 1716, 1529, 1672, 1406, 1079, 1376, 924, 1846, 1174, 1635, 1274, 1409, 1316, 1382, 1362, 1426, 1123, 1717, 1312, 1466, 1440, 1176, 1324, 1635, 1221, 1595, 1629, 3279, 1960, 1733, 2599, 1845, 1352, 3222, 1392, 1274, 1797, 2673, 1868, 2224, 1432, 1594, 2365, 2450, 2122, 1730, 2090, 2142, 1352, 1683, 2020, 1764, 1779, 1660, 2034, 1961, 2237, 1638, 2332, 1456, 1720, 1792, 2322, 2125, 1933, 1737, 1978, 1796, 1611, 2022, 2200, 1852, 2031, 1764, 1710, 1582, 1656, 1600, 1642, 2030, 1877, 2643, 2758, 2551, 2385, 2398, 2531, 2538, 2154, 2080, 1812, 2654, 2127, 1958, 1873, 1308, 1796, 1668, 2090, 1797, 1408, 1756, 1501, 2263, 1574, 1914, 1509, 1414, 1976, 1911, 1499, 1995, 1724, 2315, 2519, 1560, 1482, 1768, 1427, 1369, 1344, 2168, 2358, 1086, 2601, 1660, 1824, 1355, 1098, 1428, 2009, 1436, 1784, 2104, 1374, 1152, 1034, 1382, 1472, 1374, 1430, 2071, 1166, 1165, 1350, 1656, 954, 996, 965, 768, 1034, 898, 999, 1291),
bonus = c(0, 1, 0, 0, 1, 0, 0, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 1, 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 1, 1, 1, 0, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 1, 0, 1, 1, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 0, 1, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 0, 0, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 1, 0, 1, 1, 1, 1, 0, 1, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 1, 0, 0, 0, 1, 1, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0),
n=300
)
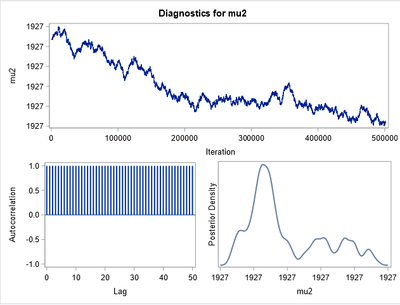
My problem is that when I try to reproduce this in Proc MCMC I get crazy results, without converge and in the autocorrelation I have zebra patterns. I think* the problem is in
model area ~ dnorm(mu[bonus+1],tau[bonus+1])
I say I think because the log is clean and all I have is my suspicion that mu[1] and tau[1] fail when bonus=1 and mu[2] and tau[2] fail when bonus=0. A failure is understood as not taking the record as null and ignoring it.
If someone has any idea, I would appreciate it.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Google search:
bayesian t-test site:communities.sas.com
Koen
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
this is, the right way to translate from BUGS to SAS/MCMC.The t-test is
just one example of the problem with different management of missing/NA in
BUGS and missing/. In SAS.
Greetings
Available on demand!
Missed SAS Innovate Las Vegas? Watch all the action for free! View the keynotes, general sessions and 22 breakouts on demand.
ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.