- Home
- /
- Analytics
- /
- Stat Procs
- /
- Linkage Clustering
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I am doing Linkage Clustering with ward's method and i noticed the Pseudo F Statistic is like huge 40000-50000, I know it's bigger is better. But is such huge number normal?
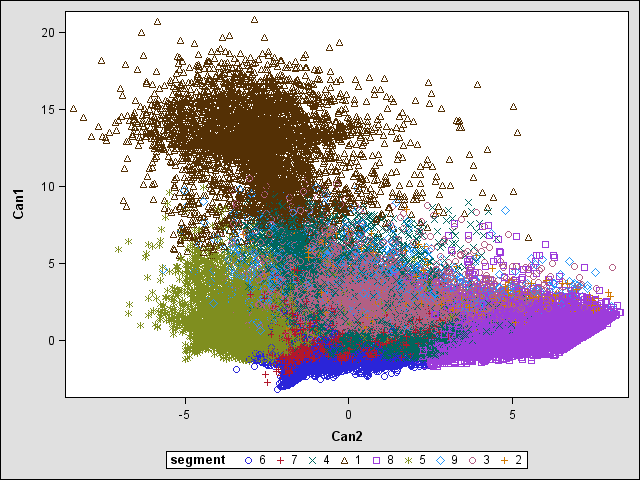
Also, i done Canonical discriminant analysis (Plot Below), my manager is questioning why some segment is so spread apart ie: segment 9, the light blue one. I explain that it's only a 2D projection of all the factors so it's normal but he's not buying it. How should i explain it to him.
proc cluster data=myData method=ward ccc pseudo;
var factor1-factor7.;
copy clus0;
run;
proc tree noprint ncl=9 out=out;
copy factor1-factor&numFactors. clus0;
run;
| Number of Clusters | Clusters Joined | Freq | Semipartial R-Square | R-Square | Approximate Expected R-Square | Cubic Clustering Criterion | Pseudo F Statistic | Pseudo t-Squared | |
| 19 | OB15 | OB18 | 736 | 0.0035 | 0.708 | 0.572 | 764 | 4.00E+04 | 460 |
| 18 | OB9 | OB14 | 3861 | 0.0049 | 0.703 | 0.566 | 772 | 4.20E+04 | 1880 |
| 17 | OB10 | CL19 | 1521 | 0.0079 | 0.695 | 0.558 | 763 | 4.30E+04 | 852 |
| 16 | OB1 | OB16 | 6658 | 0.0087 | 0.687 | 0.551 | 753 | 4.40E+04 | 3047 |
| 15 | OB4 | OB13 | 11577 | 0.0142 | 0.673 | 0.542 | 712 | 4.40E+04 | 5594 |
| 14 | CL16 | OB5 | 8143 | 0.0143 | 0.658 | 0.533 | 677 | 4.40E+04 | 3017 |
| 13 | CL18 | CL17 | 5382 | 0.0143 | 0.644 | 0.523 | 648 | 4.50E+04 | 2088 |
| 12 | OB12 | OB20 | 18770 | 0.0167 | 0.627 | 0.512 | 613 | 4.60E+04 | 1.50E+04 |
| 11 | OB8 | OB17 | 67981 | 0.0168 | 0.61 | 0.5 | 586 | 4.70E+04 | 2.50E+04 |
| 10 | CL12 | OB19 | 41771 | 0.0195 | 0.591 | 0.486 | 553 | 4.80E+04 | 1.50E+04 |
| 9 | OB7 | OB11 | 15612 | 0.0204 | 0.571 | 0.47 | 529 | 5.00E+04 | 8577 |
| 8 | CL11 | CL10 | 109752 | 0.022 | 0.549 | 0.452 | 511 | 5.20E+04 | 1.70E+04 |
| 7 | OB2 | OB6 | 125900 | 0.0523 | 0.496 | 0.425 | 346 | 4.90E+04 | 1.00E+05 |
| 6 | CL7 | CL15 | 137477 | 0.059 | 0.437 | 0.39 | 212 | 4.60E+04 | 5.10E+04 |
| 5 | CL14 | CL8 | 117895 | 0.0746 | 0.362 | 0.345 | 74.2 | 4.30E+04 | 4.10E+04 |
| 4 | CL6 | CL9 | 153089 | 0.078 | 0.284 | 0.29 | -21 | 4.00E+04 | 4.30E+04 |
| 3 | CL5 | CL4 | 270984 | 0.0813 | 0.203 | 0.216 | -57 | 3.80E+04 | 3.40E+04 |
| 2 | CL3 | OB3 | 293984 | 0.1008 | 0.102 | 0.125 | -118 | 3.40E+04 | 4.00E+04 |
| 1 | CL2 | CL13 | 299366 | 0.1023 | 0 | 0 | 0 | . | 3.40E+04 |
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The light blue segment is spread out because:
The data in the light blue segment is spread out.
Regarding the comment by @mkeintz, yes, rotating the view would certainly cause some colors to look more spread out or less spread out, but when you have 2 dimensions and you plot the results in 2 dimensions I'm not sure why rotation is needed or meaningful. But anyway, you can't get spread out data unless the data itself is spread out; if the data was really dispersed over a TINY area, no rotation would change that.
Pseudo F statistic 40000-50000 because that's what the data is saying. Perhaps it's an outlier, perhaps the clusters really really really really are distinct and separated. How can we say without your data?
Paige Miller
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
"he's not buying it?" Tell him if you rotated the data in another dimension, all those clusters looking relatively compact in this plot may very will look quite dispersed from another angle. And the one in question may look more compact.
The hash OUTPUT method will overwrite a SAS data set, but not append. That can be costly. Consider voting for Add a HASH object method which would append a hash object to an existing SAS data set
Would enabling PROC SORT to simultaneously output multiple datasets be useful? Then vote for
Allow PROC SORT to output multiple datasets
--------------------------
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
The light blue segment is spread out because:
The data in the light blue segment is spread out.
Regarding the comment by @mkeintz, yes, rotating the view would certainly cause some colors to look more spread out or less spread out, but when you have 2 dimensions and you plot the results in 2 dimensions I'm not sure why rotation is needed or meaningful. But anyway, you can't get spread out data unless the data itself is spread out; if the data was really dispersed over a TINY area, no rotation would change that.
Pseudo F statistic 40000-50000 because that's what the data is saying. Perhaps it's an outlier, perhaps the clusters really really really really are distinct and separated. How can we say without your data?
Paige Miller
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
It may be that the 2 dimension chosen is optimized to chose the largest number of clusters as compact, possibly at the expense of apparent compactness of a particular cluster.
The hash OUTPUT method will overwrite a SAS data set, but not append. That can be costly. Consider voting for Add a HASH object method which would append a hash object to an existing SAS data set
Would enabling PROC SORT to simultaneously output multiple datasets be useful? Then vote for
Allow PROC SORT to output multiple datasets
--------------------------
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
@mkeintz wrote:
It may be that the 2 dimension chosen is optimized to chose the largest number of clusters as compact, possibly at the expense of apparent compactness of a particular cluster.
Okay, but that's not what canonical correlation does.
Paige Miller
Don't miss out on SAS Innovate - Register now for the FREE Livestream!
Can't make it to Vegas? No problem! Watch our general sessions LIVE or on-demand starting April 17th. Hear from SAS execs, best-selling author Adam Grant, Hot Ones host Sean Evans, top tech journalist Kara Swisher, AI expert Cassie Kozyrkov, and the mind-blowing dance crew iLuminate! Plus, get access to over 20 breakout sessions.
ANOVA, or Analysis Of Variance, is used to compare the averages or means of two or more populations to better understand how they differ. Watch this tutorial for more.
Find more tutorials on the SAS Users YouTube channel.