- Home
- /
- Programming
- /
- SAS Procedures
- /
- ERROR: There are no individuals with any non-missing genotypes.
- RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I tried to run the HWE test using following procedure - And I got the following error
ERROR: There are no individuals with any non-missing genotypes.
One of the my observation has value Undeter instead of AA or GG etc
How can I fix this issue?
proc allele data=SNP393_A outstat=ld prefix=Marker
perms=10000 boot=1000 seed=123 genocol;
var g1-g5;
run;
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
I'm not familiar with the procedure ALLELE but the error you are getting suggests that your variables g1 to g5 contain only missing values. These variables need to be populated for at least some of your data rows to fix the error.
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
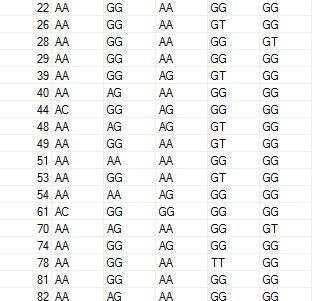
This is how my dataset looks like!
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
Read the documentation for the PROC ALLELE statement:
GENOCOL
indicates that columns specified in the VAR statement contain genotypes instead of alleles. When this option is specified, there is one column per marker. The genotypes must consist of the two alleles separated by a delimiter. For a genotype with one missing allele, use a blank space to indicate a missing value; if both alleles are missing, either use a single missing value for the entire genotype or use the delimiter alone.
(Bold and underline by me)
- Mark as New
- Bookmark
- Subscribe
- Mute
- RSS Feed
- Permalink
- Report Inappropriate Content
proc allele data=SNP393_A outstat=ld
perms=10000 boot=1000 seed=123 genocol delimiter='';
var g1-g5;
run;
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →Learn the difference between classical and Bayesian statistical approaches and see a few PROC examples to perform Bayesian analysis in this video.
Find more tutorials on the SAS Users YouTube channel.
SAS Training: Just a Click Away
Ready to level-up your skills? Choose your own adventure.




