- Home
- /
- SAS Communities Library
- /
- Finding a complete sub matrix (aka finding maximal bicliques)
- RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Finding a complete sub matrix (aka finding maximal bicliques)
- Article History
- RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Complete subsets
Here is a situation that I have met countless times. I want to explore a data matrix containing missing values with a multivariate method such as principal components, discriminant analysis, or clustering. In these and other multivarate methods, observations that have missing values for any variable are omitted from the analysis. Thus, with my full set of variables, the remaining useable observations represent only a small fraction of my data. I am left with having to choose which variables to leave out of the analysis in order to get an interesting data set. Some authors have proposed statistical properties of data subsets to help choose among them but they still need a method to generate a list of candidates (e.g. Strauss, 2006, used brute force to generate the subsets).
For example, take the following artificial data matrix, where non-missing values are represented by 1
ID A B C D E F G H I J K L
1 . . . . 1 . 1 . . . . 1
2 . . . . 1 . 1 . . . . 1
3 . . . . 1 . 1 . . . . 1
4 . . . . 1 . 1 . . . . 1
5 . . . 1 . . 1 1 . 1 . 1
6 . . . . 1 . 1 . . . . 1
7 1 1 1 . . 1 . 1 1 . 1 .
8 . . . 1 . . 1 1 . 1 . 1
9 1 1 1 . 1 1 1 . 1 . 1 1
10 1 1 1 . . 1 . . 1 . 1 .
11 . . . . 1 . 1 . . . . 1
12 1 1 1 . . 1 . . 1 . 1 .
13 . . 1 . 1 . 1 . . . . 1
14 . . . 1 . . 1 1 . 1 . 1
15 . . . 1 . . . . . . 1 .
16 . . . . 1 . 1 . . . . 1
17 1 1 1 . 1 1 1 . 1 . 1 1
18 . . . 1 . . 1 1 . 1 . 1
19 . . . 1 . . 1 1 . 1 . 1
20 . . . 1 . . 1 1 . 1 . 1
We would like to identify complete subsets in that matrix. Formally, a complete subset is a maximal group of lines and columns that contains no missing values. You may verify that the matrix can be permuted (rows and columns) to
ID K B F A I C E L G J H D
12 1 1 1 1 1 1 . . . . . .
7 1 1 1 1 1 1 . . . . 1 .
10 1 1 1 1 1 1 . . . . . .
9 1 1 1 1 1 1 1 1 1 . . .
17 1 1 1 1 1 1 1 1 1 . . .
13 . . . . . 1 1 1 1 . . .
11 . . . . . . 1 1 1 . . .
2 . . . . . . 1 1 1 . . .
1 . . . . . . 1 1 1 . . .
3 . . . . . . 1 1 1 . . .
16 . . . . . . 1 1 1 . . .
6 . . . . . . 1 1 1 . . .
4 . . . . . . 1 1 1 . . .
18 . . . . . . . 1 1 1 1 1
20 . . . . . . . 1 1 1 1 1
8 . . . . . . . 1 1 1 1 1
14 . . . . . . . 1 1 1 1 1
19 . . . . . . . 1 1 1 1 1
5 . . . . . . . 1 1 1 1 1
15 1 . . . . . . . . . . 1
where many (but not all) complete subsets appear as solid rectangles. The matrix thus contains a single complete subset of 32 values and 2 variables (L, G) as well as three complete subsets of 30 values with 6, 3 and 5 variables. Eight other smaller complete subsets also exist.
In some cases, finding an interesting subset of variables is a simple task. Just remove those variables that were not measured often enough and you are left with an exploitable data set. This is what I usually try first. But sometimes, despite my best efforts, I can't be certain that I looked at every promising subset and that I ultimately made the right choice.
My latest real life encounter with this problem involved the following data structure (16 variables, 60 obs, non-missing values replaced by 1):
data AB;
do i = 1 to 4;
input (variable_1-variable_16) (1.) +1 @;
id + 1;
output;
end;
drop i;
datalines;
.......1....1111 11111.11111.1111 11111.11111.1111 11111.11111.1111
11111.11111.11.1 11111.11111.1111 11111.11111.1111 11111.11111.1111
111.1.11111.1111 11111.11111.1111 1111111111111111 ............1111
1111111111111111 ............1111 1111111111111111 1111111111111.11
1111111111111111 1111111111111111 1111111111111111 ............1111
1111111111111111 1111111111111111 1111111111111111 1111111111111.11
1111111111111111 1111111111111111 1111111111111111 1111111111111111
....1.11.1..1111 ......11....1111 ............1111 1111111111111.11
1111111111111.11 1111111111111.11 1111111111.11111 1111111111111111
1111111111.11111 1111111111111111 1111111111111111 1111111111111111
............1111 1111111111111111 1111111111111111 1111.11111.11111
1111111111111111 1111111111111111 1111111111111111 1111.11111.11111
1111111111111111 1111.11111.11111 ............1111 1111111111111111
1111111111111111 1111111111111111 1111111111111111 1111111111111111
1111111111111111 1111111111111111 1111111111111111 11111111111111.1
;
We finally decided to run a principal component analysis on 7 variables which gave us a complete subset of 47 observations. But I kept wondering what other complete subsets we could/should have considered. Back to this example later.
Technical stuff (may be skipped)
The problem of finding all complete subsets in a data matrix is identical to the well known problem of "enumerating maximal bicliques", in graph theory. It has been shown to be impossible to accomplish this task in polynomial time (i.e., it is NP-hard) for the worst cases. Algorithms exist however to perform the enumeration of maximal bicliques for moderate size graphs.
In simple terms, a biclique is a subgraph composed of two categories of nodes where every node of the first category is linked to every node of the other category. In data matrix terms, the first node category represents observations and nodes of the second category are variables. The links between variables and observations represent the presence of non-missing values in the matrix. Finding a biclique in such a graph is thus the same as identifying a complete subset in a matrix.
SAS doesn't offer a procedure for enumerating maximal bicliques, but SAS/OR does have a tool for finding all maximal cliques in undirected graphs. The OPTNET procedure has the CLIQUE statement that implements a variant of the Bron-Kerbosch algorithm. The documentation includes a warning about the NP-hardness of the problem and the CLIQUE statement provides option MAXTIME= to limit execution times.
Cliques are simpler than bicliques. They are subgraphs where every node is linked to all other nodes. A maximal clique is a clique that isn't a subgraph of any other clique.
It turns out that we can find all maximal bicliques by enumerating maximal cliques for specially devised graphs. The graph to be solved has the following structure. Every variable and every observation is a node. There are links between every pair of variables. There are also links between every pair of observations. Finally, there are links between variables and observations where the variable value is not missing. The resulting maximal cliques in that graph are sets of variables and observations that constitute maximal subsets. In some cases the set of all observations or the set of all variables may be identified as maximal cliques and should be ignored.
The completeSubsets macro
The completeSubsets macro builds a graph representing the non missing data in the input matrix and calls PROC OPTNET (requires SAS/OR license) to get the maximal cliques. The cliques are then reinterpreted for output as complete subsets.
The macro is called like this:
%completeSubsets(inDsn, outDsn, id=id, vars=_numeric_, maxTime=100);
- Where
inDsn : input data set name
outDsn : output data set name
id : name of a variable in the input data set that identifies observations
vars : list of numeric variables in input data set where subsets are to be searched
maxTime : time limit (in seconds) for the search
The input data set is your data matrix in the same format that you would submit to a SAS multivariate procedure. Note that a matrix and its transposed version will yield essentially the same subsets. The output data set will contain three variables:
- subset: a number identifying the subset
type: "OBS" or "VAR"
name: the name of a variable (if type="VAR") or the identifier of an observation (if type="OBS") as a character string.
Real-life example
For my matrix AB above, I would call
%completeSubsets(AB, mySubsets, id=id, vars=variable_1-variable_16);
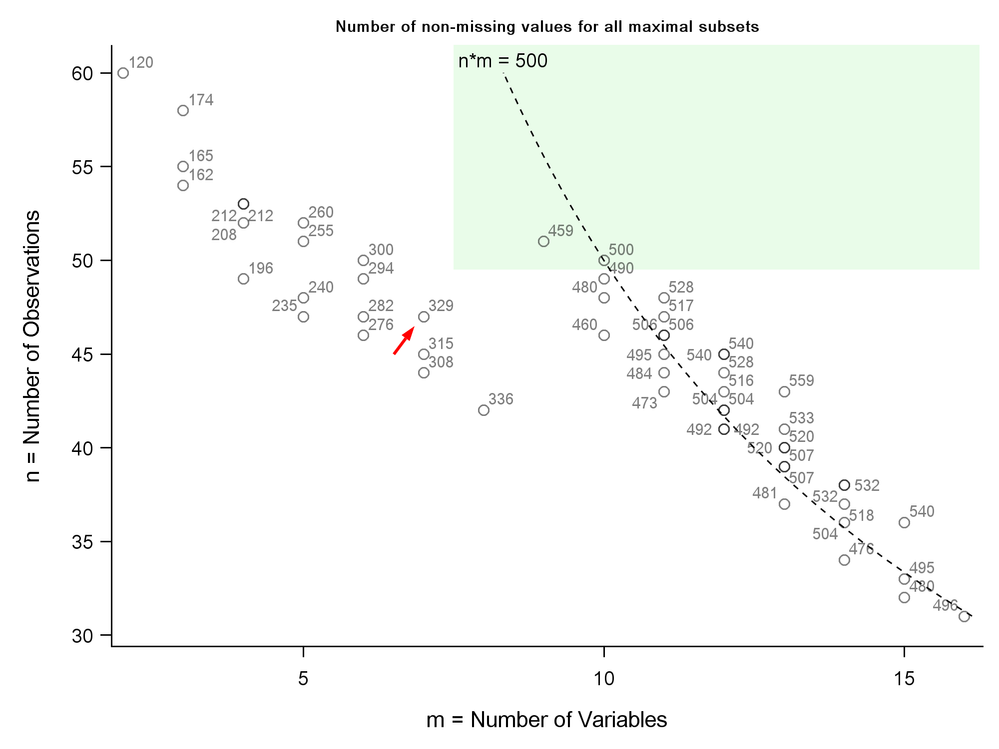
Computation of the subsets is surprisingly fast. The output data set mySubsets contains 3046 observations describing 56 subsets. Now a bit of post processing (code not shown) can rank the subsets on various properties and narrow down our choices. If matrix size for example had been the criterion for choosing a subset (it wasn't), our 7 by 47 subset from PCA (arrow) would have been shown to be rather poor...
References
Peeters, R. (2003). The maximum edge biclique problem is NP-complete. Discrete Applied Mathematics, 131:651-654.
Strauss, R.E., Atanassov, M.N. (2006) Determining best complete subsets of specimens and characters for multivariate morphometric studies in the presence of large amounts of missing data. Biological Journal of the Linnean Society, 88:309-328.
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Hmm. Very interesting question. Of course, I would not miss such kind of question.
Using GA in IML with SAS University Edition. Not SAS/OR .
data AB;
do i = 1 to 4;
input (variable_1-variable_16) (1.) +1 @;
id + 1;
output;
end;
drop i;
datalines;
.......1....1111 11111.11111.1111 11111.11111.1111 11111.11111.1111
11111.11111.11.1 11111.11111.1111 11111.11111.1111 11111.11111.1111
111.1.11111.1111 11111.11111.1111 1111111111111111 ............1111
1111111111111111 ............1111 1111111111111111 1111111111111.11
1111111111111111 1111111111111111 1111111111111111 ............1111
1111111111111111 1111111111111111 1111111111111111 1111111111111.11
1111111111111111 1111111111111111 1111111111111111 1111111111111111
....1.11.1..1111 ......11....1111 ............1111 1111111111111.11
1111111111111.11 1111111111111.11 1111111111.11111 1111111111111111
1111111111.11111 1111111111111111 1111111111111111 1111111111111111
............1111 1111111111111111 1111111111111111 1111.11111.11111
1111111111111111 1111111111111111 1111111111111111 1111.11111.11111
1111111111111111 1111.11111.11111 ............1111 1111111111111111
1111111111111111 1111111111111111 1111111111111111 1111111111111111
1111111111111111 1111111111111111 1111111111111111 11111111111111.1
;
run;
proc iml;
use AB(drop=id);
read all var _num_ into xx[c=vname] ;
close ;
start function(x) global(nrow,total,xx);
idx_row=loc(x[1:nrow]);
idx_col=loc(x[nrow+1:total]);
sum=sum(x);
if all(xx[idx_row,idx_col])=1 then obj=sum;
else obj=0;
return (obj);
finish;
nrow=nrow(xx);
ncol=ncol(xx);
total=nrow+ncol;
encoding=j(2,total,0);
encoding[2,]=1;
id=gasetup(2,total,123456789);
call gasetobj(id,1,"function");
call gasetsel(id,100,1,1);
call gainit(id,1000,encoding);
niter = 100000;
do i = 1 to niter;
call garegen(id);
call gagetval(value, id);
end;
call gagetmem(mem, value, id, 1);
idx_row=loc(mem[1:nrow]);
idx_col=loc(mem[nrow+1:total]);
names=vname[idx_col];
row=char(idx_row);
want=xx[idx_row,idx_col];
print want[l='Max Sub-Matrix' r=row c=names] ,
value[l = "Max Value:"] ;
call gaend(id);
quit;
OUTPUT:
Max Sub-Matrix
variable_1 variable_2 variable_4 variable_7 variable_8 variable_9 variable_10 variable_13 variable_16
2 1 1 1 1 1 1 1 1 1
3 1 1 1 1 1 1 1 1 1
4 1 1 1 1 1 1 1 1 1
5 1 1 1 1 1 1 1 1 1
6 1 1 1 1 1 1 1 1 1
7 1 1 1 1 1 1 1 1 1
8 1 1 1 1 1 1 1 1 1
10 1 1 1 1 1 1 1 1 1
11 1 1 1 1 1 1 1 1 1
13 1 1 1 1 1 1 1 1 1
15 1 1 1 1 1 1 1 1 1
16 1 1 1 1 1 1 1 1 1
17 1 1 1 1 1 1 1 1 1
18 1 1 1 1 1 1 1 1 1
19 1 1 1 1 1 1 1 1 1
21 1 1 1 1 1 1 1 1 1
22 1 1 1 1 1 1 1 1 1
23 1 1 1 1 1 1 1 1 1
24 1 1 1 1 1 1 1 1 1
25 1 1 1 1 1 1 1 1 1
26 1 1 1 1 1 1 1 1 1
27 1 1 1 1 1 1 1 1 1
28 1 1 1 1 1 1 1 1 1
32 1 1 1 1 1 1 1 1 1
33 1 1 1 1 1 1 1 1 1
34 1 1 1 1 1 1 1 1 1
35 1 1 1 1 1 1 1 1 1
36 1 1 1 1 1 1 1 1 1
37 1 1 1 1 1 1 1 1 1
39 1 1 1 1 1 1 1 1 1
40 1 1 1 1 1 1 1 1 1
42 1 1 1 1 1 1 1 1 1
43 1 1 1 1 1 1 1 1 1
44 1 1 1 1 1 1 1 1 1
45 1 1 1 1 1 1 1 1 1
46 1 1 1 1 1 1 1 1 1
47 1 1 1 1 1 1 1 1 1
48 1 1 1 1 1 1 1 1 1
49 1 1 1 1 1 1 1 1 1
50 1 1 1 1 1 1 1 1 1
52 1 1 1 1 1 1 1 1 1
53 1 1 1 1 1 1 1 1 1
54 1 1 1 1 1 1 1 1 1
55 1 1 1 1 1 1 1 1 1
57 1 1 1 1 1 1 1 1 1
59 1 1 1 1 1 1 1 1 1
60 1 1 1 1 1 1 1 1 1
Max Value:
56
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
The following code is more robust .
data AB;
infile cards expandtabs truncover;
input ID A B C D E F G H I J K L;
cards;
1 . . . . 1 . 1 . . . . 1
2 . . . . 1 . 1 . . . . 1
3 . . . . 1 . 1 . . . . 1
4 . . . . 1 . 1 . . . . 1
5 . . . 1 . . 1 1 . 1 . 1
6 . . . . 1 . 1 . . . . 1
7 1 1 1 . . 1 . 1 1 . 1 .
8 . . . 1 . . 1 1 . 1 . 1
9 1 1 1 . 1 1 1 . 1 . 1 1
10 1 1 1 . . 1 . . 1 . 1 .
11 . . . . 1 . 1 . . . . 1
12 1 1 1 . . 1 . . 1 . 1 .
13 . . 1 . 1 . 1 . . . . 1
14 . . . 1 . . 1 1 . 1 . 1
15 . . . 1 . . . . . . 1 .
16 . . . . 1 . 1 . . . . 1
17 1 1 1 . 1 1 1 . 1 . 1 1
18 . . . 1 . . 1 1 . 1 . 1
19 . . . 1 . . 1 1 . 1 . 1
20 . . . 1 . . 1 1 . 1 . 1
;
run;
proc iml;
use AB(drop=id);
read all var _num_ into xx[c=vname] ;
close ;
start function(x) global(nrow,total,xx);
temp=x[1:nrow];
if all(temp=0) then idx_row=1;
else idx_row=loc(temp);
temp=x[nrow+1:total];
if all(temp=0) then idx_col=1;
else idx_col=loc(temp);
sum=sum(x);
if all(xx[idx_row,idx_col])=1 then obj=sum;
else obj=0;
return (obj);
finish;
nrow=nrow(xx);
ncol=ncol(xx);
total=nrow+ncol;
encoding=j(2,total,0);
encoding[2,]=1;
id=gasetup(2,total,123456789);
call gasetobj(id,1,"function");
call gasetsel(id,100,1,1);
call gainit(id,1000,encoding);
niter = 100000;
do i = 1 to niter;
call garegen(id);
call gagetval(value, id);
end;
call gagetmem(mem, value, id, 1);
idx_row=loc(mem[1:nrow]);
idx_col=loc(mem[nrow+1:total]);
names=vname[idx_col];
row=char(idx_row);
want=xx[idx_row,idx_col];
print want[l='Max Sub-Matrix' r=row c=names] ,
value[l = "Max Value:"] ;
call gaend(id);
quit;
OUTPUT:
Max Sub-Matrix
G L
1 1 1
2 1 1
3 1 1
4 1 1
5 1 1
6 1 1
8 1 1
9 1 1
11 1 1
13 1 1
14 1 1
16 1 1
17 1 1
18 1 1
19 1 1
20 1 1
Max Value:
18
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Great! You demonstrate once again that GA are very versatile. But the network algorithm in PROC OPTNET finds ALL sub-matrices for both examples in less than 0.1 s CPU time!
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
There is this warning in the PROC OPTNET - CLIQUE documentation
The algorithm that PROC OPTNET uses to compute maximal cliques is a variant of the
Bron-Kerbosch algorithm (Bron and Kerbosch, 1973; Harley, 2003). Enumerating all
maximal cliques is NP-hard, so this algorithm typically does not scale to very large graphs.
Note: the warning is about the intrinsic difficulty of finding ALL maximal cliques.
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Max Sub-Matrix variable_13 variable_16 1 1 1 2 1 1 3 1 1 4 1 1 5 1 1 6 1 1 7 1 1 8 1 1 9 1 1 10 1 1 11 1 1 12 1 1 13 1 1 14 1 1 15 1 1 16 1 1 17 1 1 18 1 1 19 1 1 20 1 1 21 1 1 22 1 1 23 1 1 24 1 1 25 1 1 26 1 1 27 1 1 28 1 1 29 1 1 30 1 1 31 1 1 32 1 1 33 1 1 34 1 1 35 1 1 36 1 1 37 1 1 38 1 1 39 1 1 40 1 1 41 1 1 42 1 1 43 1 1 44 1 1 45 1 1 46 1 1 47 1 1 48 1 1 49 1 1 50 1 1 51 1 1 52 1 1 53 1 1 54 1 1 55 1 1 56 1 1 57 1 1 58 1 1 59 1 1 60 1 1 62
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Yes, maximum of m=2 variables + n=60 (all) observations. It is the point on the upper left of the Figure. Others may want to maximize n or m or n*m, with possibly some constraints such as m>=8 and n>=50 (green rectangle), or n*m>500 (on the right of the dashed line), etc.
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Here's how to maximize n * m by using the MILP solver in SAS/OR:
data orig_links;
array variable_[16];
set AB;
from = 'row'||put(id,z2.);
do j = 1 to 16;
if variable_[j] then do;
to = 'col'||put(j,z2.);
output;
end;
end;
run;
proc optmodel;
/* read in links from bipartite graph */
set <str,str> LINKS;
read data orig_links into LINKS=[from to];
set <str> LEFT_NODES init {};
set <str> RIGHT_NODES init {};
for {<i,j> in LINKS} do;
LEFT_NODES = LEFT_NODES union {i};
RIGHT_NODES = RIGHT_NODES union {j};
end;
put (card(LEFT_NODES))= (card(RIGHT_NODES))=;
set NODES = LEFT_NODES union RIGHT_NODES;
/* UseNode[i] = 1 if node i appears in biclique, 0 otherwise */
var UseNode {NODES} binary;
/* UseLink[i,j] = 1 if link <i,j> appears in biclique, 0 otherwise */
var UseLink {LINKS} binary;
/* maximize number of edges */
max NumEdges = sum {<i,j> in LINKS} UseLink[i,j];
/* if <i,j> not in LINKS, cannot have both nodes i and j */
con Conflict {i in LEFT_NODES, j in RIGHT_NODES: <i,j> not in LINKS}:
UseNode[i] + UseNode[j] <= 1;
/* if UseLink[i,j] = 1 then UseNode[i] = 1 and UseNode[j] = 1 */
con Link_i {<i,j> in LINKS}:
UseLink[i,j] <= UseNode[i];
con Link_j {<i,j> in LINKS}:
UseLink[i,j] <= UseNode[j];
solve;
set NODES_SOL = {i in NODES: UseNode[i].sol > 0.5};
put (card(NODES_SOL inter LEFT_NODES))= (card(NODES_SOL inter RIGHT_NODES))=;
set LINKS_SOL = {i in NODES_SOL, j in NODES_SOL} inter LINKS;
create data maxbiclique_nodes from [node]=NODES_SOL;
create data maxbiclique_links from [from to]=LINKS_SOL;
quit;- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Mine is maximize n+m . Of course , it is easy to maximize n or m or n*m ........ for GA . GA almost can solve all kind of OR problems.That attract me . I have no time and SAS/OR to learn its language . The reason might be you need pay it .
Catch up on SAS Innovate 2026
Nearly 200 sessions are now available on demand with the SAS Innovate Digital Pass.
Explore Now →SAS AI and Machine Learning Courses
The rapid growth of AI technologies is driving an AI skills gap and demand for AI talent. Ready to grow your AI literacy? SAS offers free ways to get started for beginners, business leaders, and analytics professionals of all skill levels. Your future self will thank you.